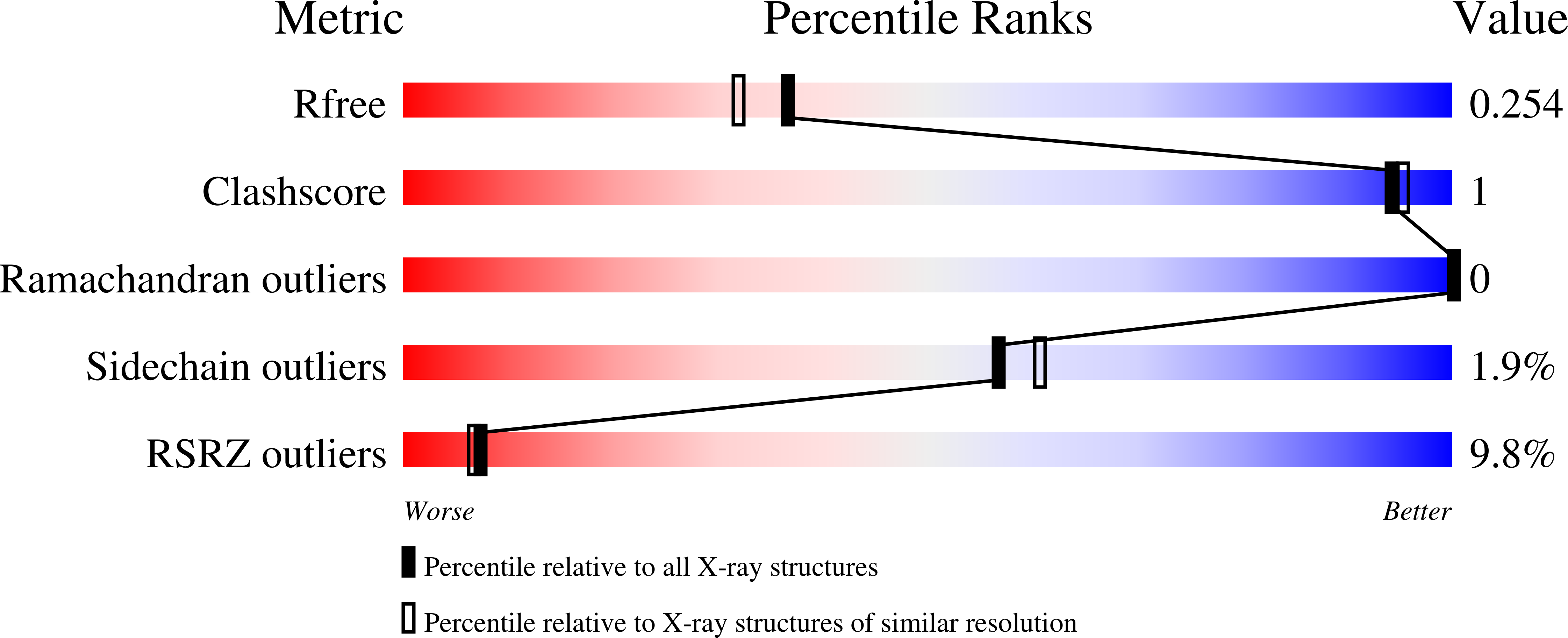
Characterization of a trifunctional mimivirus mRNA capping enzyme and crystal structure of the RNA triphosphatase domain.
Benarroch, D., Smith, P., Shuman, S.(2008) Structure 16: 501-512
- PubMed: 18400173
- DOI: https://doi.org/10.1016/j.str.2008.01.009
- Primary Citation of Related Structures:
2QY2, 2QZE - PubMed Abstract:
The RNA triphosphatase (RTPase) components of the mRNA capping apparatus are a bellwether of eukaryal taxonomy. Fungal and protozoal RTPases belong to the triphosphate tunnel metalloenzyme (TTM) family, exemplified by yeast Cet1. Several large DNA viruses encode metal-dependent RTPases unrelated to the cysteinyl-phosphatase RTPases of their metazoan host organisms. The origins of DNA virus RTPases are unclear because they are structurally uncharacterized. Mimivirus, a giant virus of amoeba, resembles poxviruses in having a trifunctional capping enzyme composed of a metal-dependent RTPase module fused to guanylyltransferase (GTase) and guanine-N7 methyltransferase domains. The crystal structure of mimivirus RTPase reveals a minimized tunnel fold and an active site strikingly similar to that of Cet1. Unlike homodimeric fungal RTPases, mimivirus RTPase is a monomer. The mimivirus TTM-type RTPase-GTase fusion resembles the capping enzymes of amoebae, providing evidence that the ancestral large DNA virus acquired its capping enzyme from a unicellular host.
Organizational Affiliation:
Molecular Biology Program, Sloan-Kettering Institute, New York, NY 10021, USA.