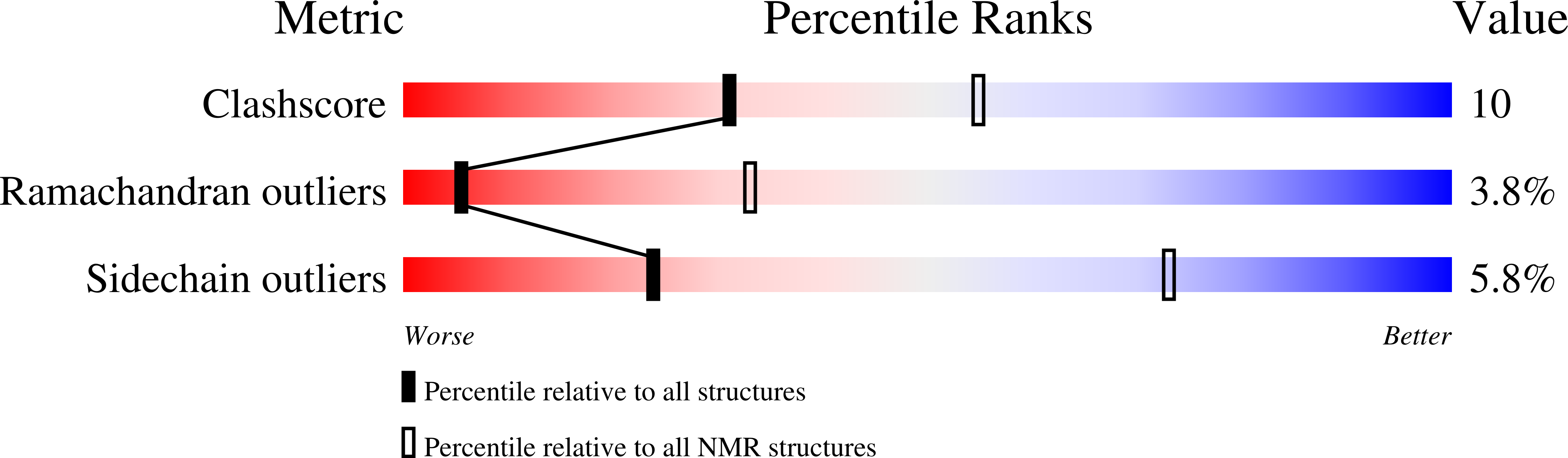
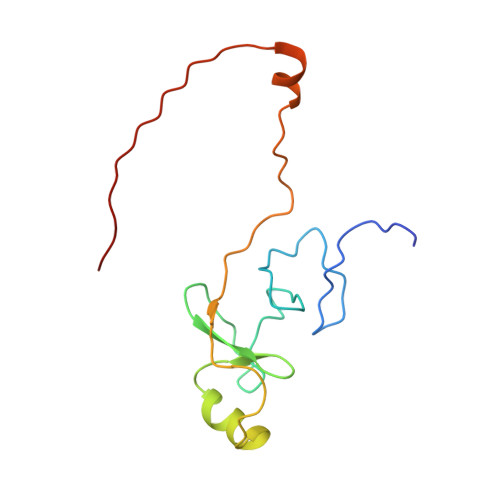
Solution structure of a tethered Lmo2(LIM2) /Ldb1(LID) complex.
Dastmalchi, S., Wilkinson-White, L., Kwan, A.H., Gamsjaeger, R., Mackay, J.P., Matthews, J.M.(2012) Protein Sci 21: 1768-1774
- PubMed: 22936624
- DOI: https://doi.org/10.1002/pro.2153
- Primary Citation of Related Structures:
2LXD - PubMed Abstract:
LIM-only protein 2, Lmo2, is a regulatory protein that is essential for hematopoietic development and inappropriate overexpression of Lmo2 in T-cells contributes to T-cell leukemia. It exerts its functions by mediating protein-protein interactions and nucleating multicomponent transcriptional complexes. Lmo2 interacts with LIM domain binding protein 1 (Ldb1) through the tandem LIM domains of Lmo2 and the LIM interaction domain (LID) of Ldb1. Here, we present the solution structure of the LIM2 domain of Lmo2 bound to Ldb1(LID) . The ordered regions of Ldb1 in this complex correspond well with binding hotspots previously defined by mutagenic studies. Comparisons of this Lmo2(LIM2) -Ldb1(LID) structure with previously determined structures of the Lmo2/Ldb1(LID) complexes lead to the conclusion that modular binding of tandem LIM domains in Lmo2 to tandem linear motifs in Ldb1 is accompanied by several disorder-to-order transitions and/or conformational changes in both proteins.
Organizational Affiliation:
School of Molecular Bioscience, University of Sydney, Sydney, New South Wales 2006, Australia.