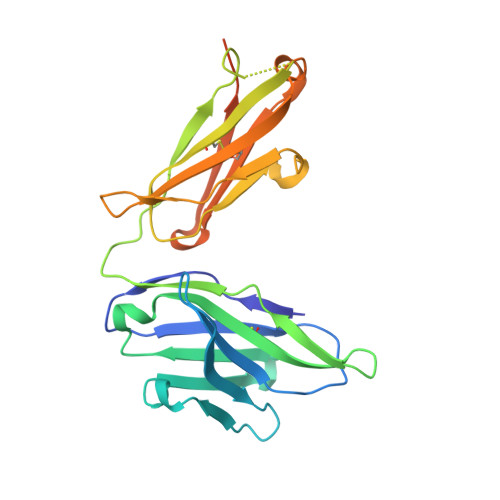
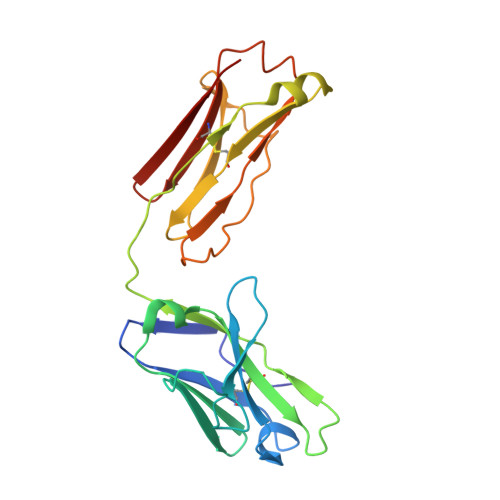
Fab Mor03268 Triggers Absorption Shift of a Diagnostic Dye Via Packaging in a Solvent-Shielded Fab Dimer Interface
Hillig, R.C., Urlinger, S., Fanghanel, J., Brocks, B., Haenel, C., Stark, Y., Sulzle, D., Svergun, D.I., Baesler, S., Malawski, G., Moosmayer, D., Menrad, A., Schirner, M., Licha, K.(2008) J Mol Biol 377: 206
- PubMed: 18241888
- DOI: https://doi.org/10.1016/j.jmb.2007.12.071
- Primary Citation of Related Structures:
2JB5, 2JB6 - PubMed Abstract:
Molecular interactions between near-IR fluorescent probes and specific antibodies may be exploited to generate novel smart probes for diagnostic imaging. Using a new phage display technology, we developed such antibody Fab fragments with subnanomolar binding affinity for tetrasulfocyanine, a near-IR in vivo imaging agent. Unexpectedly, some Fabs induced redshifts of the dye absorption peak of up to 44 nm. This is the largest shift reported for a biological system so far. Crystal structure determination and absorption spectroscopy in the crystal in combination with microcalorimetry and small-angle X-ray scattering in solution revealed that the redshift is triggered by formation of a Fab dimer, with tetrasulfocyanine being buried in a fully closed protein cavity within the dimer interface. The derived principle of shifting the absorption peak of a symmetric dye via packaging within a Fab dimer interface may be transferred to other diagnostic fluorophores, opening the way towards smart imaging probes that change their wavelength upon interaction with an antibody.
Organizational Affiliation:
Bayer Schering Pharma AG, Global Drug Discovery, 13342 Berlin, Germany. roman.hillig@bayerhealthcare.com