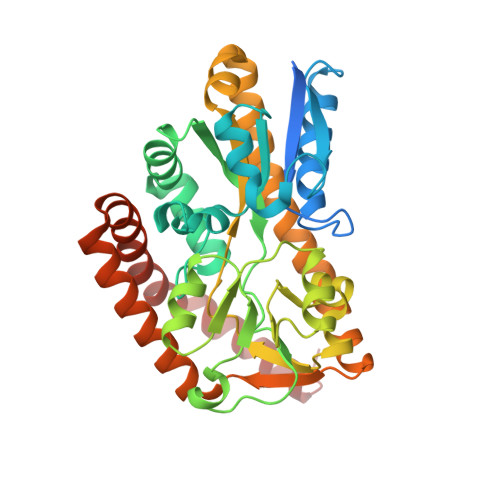
Crystal structures of an Extracytoplasmic Solute Receptor from a TRAP transporter in its open and closed forms reveal a helix-swapped dimer requiring a cation for alpha-keto acid binding.
Gonin, S., Arnoux, P., Pierru, B., Lavergne, J., Alonso, B., Sabaty, M., Pignol, D.(2007) BMC Struct Biol 7: 11-11
- PubMed: 17362499
- DOI: https://doi.org/10.1186/1472-6807-7-11
- Primary Citation of Related Structures:
2HZK, 2HZL - PubMed Abstract:
The import of solutes into the bacterial cytoplasm involves several types of membrane transporters, which may be driven by ATP hydrolysis (ABC transporters) or by an ion or H+ electrochemical membrane potential, as in the tripartite ATP-independent periplasmic system (TRAP). In both the ABC and TRAP systems, a specific periplasmic protein from the ESR family (Extracytoplasmic Solute Receptors) is often involved for the recruitment of the solute and its presentation to the membrane complex. In Rhodobacter sphaeroides, TakP (previously named SmoM) is an ESR from a TRAP transporter and binds alpha-keto acids in vitro. We describe the high-resolution crystal structures of TakP in its unliganded form and as a complex with sodium-pyruvate. The results show a limited "Venus flytrap" conformational change induced by substrate binding. In the liganded structure, a cation (most probably a sodium ion) is present and plays a key role in the association of the pyruvate to the protein. The structure of the binding pocket gives a rationale for the relative affinities of various ligands that were tested from a fluorescence assay. The protein appears to be dimeric in solution and in the crystals, with a helix-swapping structure largely participating in the dimer formation. A 30 A-long water channel buried at the dimer interface connects the two ligand binding cavities of the dimer. The concerted recruitment by TakP of the substrate group with a cation could represent a first step in the coupled transport of both partners, providing the driving force for solute import. Furthermore, the unexpected dimeric structure of TakP suggests a molecular mechanism of solute uptake by the dimeric ESR via a channel that connects the binding sites of the two monomers.
Organizational Affiliation:
CEA/Cadarache, DSV/DEVM, Laboratoire de Bioénergétique Cellulaire, 13108 St Paul lez Durance Cedex, France. sophie.gonin@cea.fr