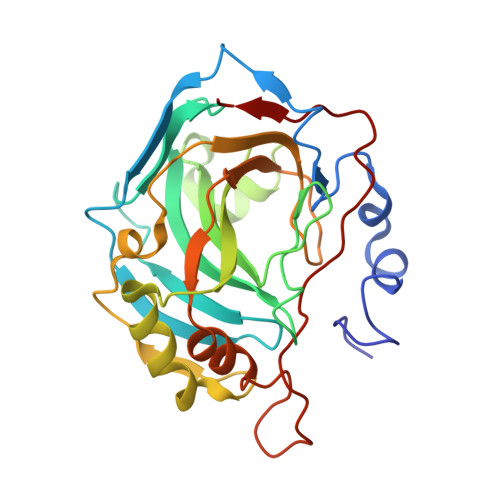
Carbonic anhydrase inhibitors: Hypoxia-activatable sulfonamides incorporating disulfide bonds that target the tumor-associated isoform IX.
De Simone, G., Vitale, R.M., Di Fiore, A., Pedone, C., Scozzafava, A., Montero, J.L., Winum, J.Y., Supuran, C.T.(2006) J Med Chem 49: 5544-5551
- PubMed: 16942027
- DOI: https://doi.org/10.1021/jm060531j
- Primary Citation of Related Structures:
2HD6 - PubMed Abstract:
An approach for designing bioreductive, hypoxia-activatable carbonic anhydrase (CA, EC 4.2.1.1) inhibitors targeting the tumor-associated isoforms is reported. Sulfonamides incorporating 3,3'-dithiodipropionamide/2,2'-dithiodibenzamido moieties were prepared and reduced enzymatically/chemically in conditions present in hypoxic tumors, leading to thiols. The X-ray crystal structure of the most promising compound, 4-(2-mercaptophenylcarboxamido)benzenesulfonamide, which as disulfide showed a K(I) against hCA IX of 653 nM (in reduced form of 9.1 nM), in adduct with hCA II showed the inhibitor making favorable interactions with Gln92, Val121, Phe131, Leu198, Thr199, Thr200, Pro201, and Pro202, whereas the sulfamoyl moiety was coordinated to the Zn2+ ion. The same interactions were preserved in the adduct with hCA IX, but in addition, a hydrogen bond between the SH moiety of the inhibitor and the amide nitrogen of Gln67 was evidenced, which may explain the almost 2 times more effective inhibition of the tumor-associated isozyme over the cytosolic isoform.
Organizational Affiliation:
Istituto di Biostrutture e Bioimmagini-CNR, Via Mezzocannone 16, 80134 Naples, Italy. gdesimone@unina.it