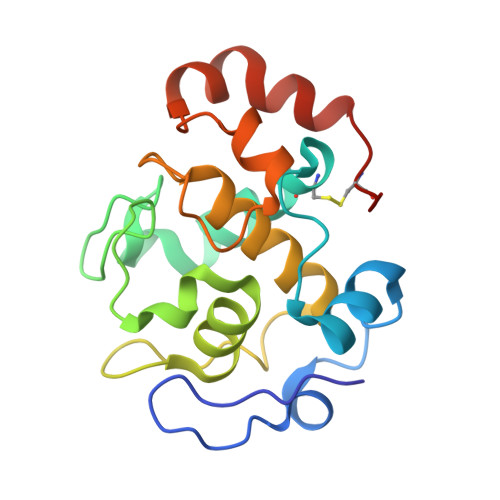
Crystal structures of cytochrome c(L) and methanol dehydrogenase from Hyphomicrobium denitrificans: structural and mechanistic insights into interactions between the two proteins
Nojiri, M., Hira, D., Yamaguchi, K., Okajima, T., Tanizawa, K., Suzuki, S.(2006) Biochemistry 45: 3481-3492
- PubMed: 16533029
- DOI: https://doi.org/10.1021/bi051877j
- Primary Citation of Related Structures:
2D0V, 2D0W - PubMed Abstract:
Methanol dehydrogenase (Hd-MDH) and its physiological electron acceptor, cytochrome c(L) (Hd-Cyt c(L)), isolated from a methylotrophic denitrifying bacterium, Hyphomicrobium denitrificans A3151, have been kinetically and structurally characterized; the X-ray structures of Hd-MDH and Hd-Cyt c(L) have been determined using molecular replacement at 2.5 and 2.0 A resolution, respectively. To explain the mechanism for electron transfer between these proteins, the dependence of MDH activity on the concentration of Hd-Cyt c(L) has been investigated at pH 4.5-7.0. The Michaelis constant for Hd-Cyt c(L) shows the smallest value (approximately 0.3 microM) at pH 5.5. The pseudo-first-order rate constant (k(obs)) of the reduction of Hd-Cyt c(L) exhibits a hyperbolic concentration dependence of Hd-MDH at pH 5.5, although k(obs) linearly increases at pH 6.5. These findings indicate formation of a transient complex between these proteins during an electron transfer event. Hd-MDH (148 kDa) is a large tetrameric protein with an alpha(2)beta(2) subunit composition, showing a high degree of structural similarity with other MDHs. Hd-Cyt c(L) (19 kDa) exhibiting the alpha-band at 550.7 nm has a unique C-terminal region involving a disulfide bond between Cys47 and Cys165. Moreover, there is a pair of Hd-Cyt c(L) monomers related with a pseudo-2-fold axis of symmetry in the asymmetric unit, and the two monomers tightly interact with each other through three hydrogen bonds. This configuration is the first example in the studies of cytochrome c as the physiological electron acceptor for MDH. The docking simulation between the coupled Hd-Cyt c(L) molecules and the heterotetrameric Hd-MDH molecule has been carried out.
Organizational Affiliation:
Department of Chemistry, Graduate School of Science, Osaka University, Toyonaka, Osaka 560-0043, Japan. nojiri@ch.wani.osaka-u.ac.jp