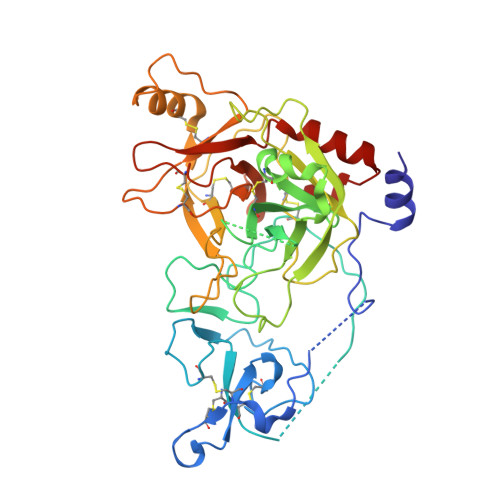
Crystal structure of a clip-domain serine protease and functional roles of the clip domains.
Piao, S., Song, Y.L., Kim, J.H., Park, S.Y., Park, J.W., Lee, B.L., Oh, B.H., Ha, N.C.(2005) EMBO J 24: 4404-4414
- PubMed: 16362048
- DOI: https://doi.org/10.1038/sj.emboj.7600891
- Primary Citation of Related Structures:
2B9L - PubMed Abstract:
Clip-domain serine proteases (SPs) are the essential components of extracellular signaling cascades in various biological processes, especially in embryonic development and the innate immune responses of invertebrates. They consist of a chymotrypsin-like SP domain and one or two clip domains at the N-terminus. Prophenoloxidase-activating factor (PPAF)-II, which belongs to the noncatalytic clip-domain SP family, is indispensable for the generation of the active phenoloxidase leading to melanization, a major defense mechanism of insects. Here, the crystal structure of PPAF-II reveals that the clip domain adopts a novel fold containing a central cleft, which is distinct from the structures of defensins with a similar arrangement of cysteine residues. Ensuing studies demonstrated that PPAF-II forms a homo-oligomer upon cleavage by the upstream protease and that the clip domain of PPAF-II functions as a module for binding phenoloxidase through the central cleft, while the clip domain of a catalytically active easter-type SP plays an essential role in the rapid activation of its protease domain.
Organizational Affiliation:
National Research Laboratory of Defense Proteins, College of Pharmacy and Research Institute for Drug Development, Pusan National University, Jangjeon Dong, Geumjeong Gu, Busan, Korea.