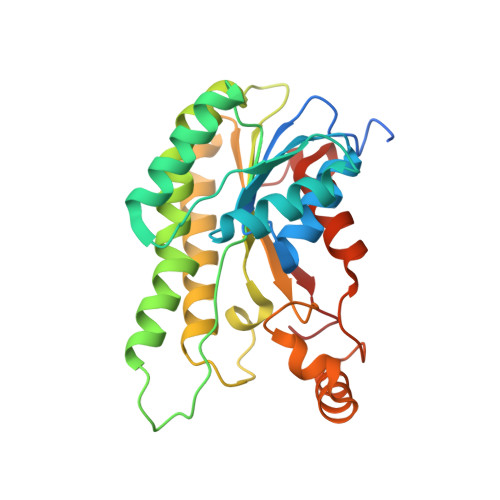
Structure of RhlG, an Essential beta-Ketoacyl Reductase in the Rhamnolipid Biosynthetic Pathway of Pseudomonas aeruginosa.
Miller, D.J., Zhang, Y.-M., Rock, C.O., White, S.W.(2006) J Biol Chem 281: 18025-18032
- PubMed: 16624803
- DOI: https://doi.org/10.1074/jbc.M601687200
- Primary Citation of Related Structures:
2B4Q - PubMed Abstract:
Rhamnolipids are extracellular biosurfactants and virulence factors secreted by the opportunistic human pathogen Pseudomonas aeruginosa that are required for swarming motility. The rhlG gene is essential for rhamnolipid formation, and the RhlG enzyme is thought to divert fatty acid synthesis intermediates into the rhamnolipid biosynthetic pathway based on its similarity to FabG, the beta-ketoacyl-acyl carrier protein (ACP) reductase of type II fatty acid synthesis. Crystallographic analysis reveals that the overall structures of the RhlG.NADP+ and FabG.NADP+ complexes are indeed similar, but there are key differences related to function. RhlG does not undergo the conformational changes upon NADP(H) binding at the active site that in FabG are the structural basis of negative allostery. Also, the acyl chain-binding pocket of RhlG is narrow and rigid compared with the larger, flexible substrate-binding subdomain in FabG. Finally, RhlG lacks a positively charged/hydrophobic surface feature adjacent to the active site that is found on enzymes like FabG that recognize the ACP of fatty acid synthesis. RhlG catalyzed the NADPH-dependent reduction of beta-ketodecanoyl-ACP to beta-d-hydroxydecanoyl-ACP. However, the enzyme was 2000-fold less active than FabG in carrying out the same reaction. These structural and biochemical studies establish RhlG as a NADPH-dependent beta-ketoacyl reductase of the SDR protein superfamily and further suggest that the ACP of fatty acid synthesis does not carry the substrates for RhlG.
Organizational Affiliation:
Department of Structural Biology, St. Jude Children's Research Hospital, 332 N. Lauderdale Street, Memphis, TN 38105, USA.