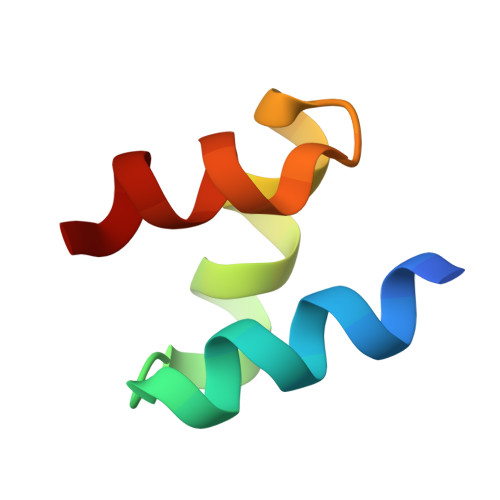
Mechanism of Lys48-linked polyubiquitin chain recognition by the Mud1 UBA domain
Trempe, J.-F., Brown, N.R., Lowe, E.D., Gordon, C., Campbell, I.D., Noble, M.E.M., Endicott, J.A.(2005) EMBO J 24: 3178-3189
- PubMed: 16138082
- DOI: https://doi.org/10.1038/sj.emboj.7600797
- Primary Citation of Related Structures:
1Z96 - PubMed Abstract:
The ubiquitin-pathway associated (UBA) domain is a 40-residue polyubiquitin-binding motif. The Schizosaccharomyces pombe protein Mud1 is an ortholog of the Saccharomyces cerevisiae DNA-damage response protein Ddi1 and binds to K48-linked polyubiquitin through its UBA domain. We have solved the crystal structure of Mud1 UBA at 1.8 angstroms resolution, revealing a canonical three-helical UBA fold. We have probed the interactions of this domain using mutagenesis, surface plasmon resonance, NMR and analytical ultracentrifugation. We show that the ubiquitin-binding surface of Mud1 UBA extends beyond previously recognized motifs and can be functionally dissected into primary and secondary ubiquitin-binding sites. Mutation of Phe330 to alanine, a residue exposed between helices 2 and 3, significantly reduces the affinity of the Mud1 UBA domain for K48-linked polyubiquitin, despite leaving the primary binding surface functionally intact. Moreover, K48-linked diubiquitin binds a single Mud1 UBA domain even in the presence of excess UBA. We therefore propose a mechanism for the recognition of K48-linked polyubiquitin chains by Mud1 in which diubiquitin units are specifically recognized by a single UBA domain.
Organizational Affiliation:
Laboratory of Molecular Biophysics, University of Oxford, Oxford, UK.