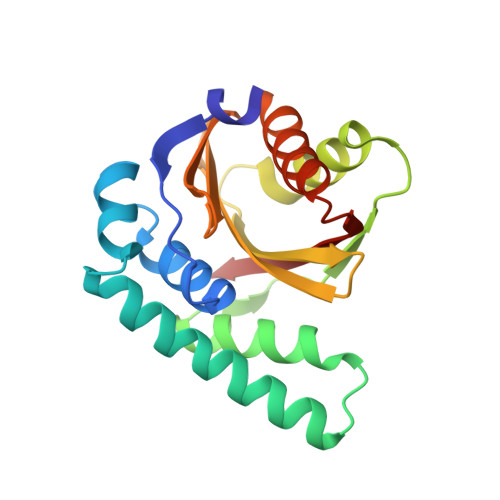
The crystal structure of Pseudomonas avirulence protein AvrPphB: a papain-like fold with a distinct substrate-binding site.
Zhu, M., Shao, F., Innes, R.W., Dixon, J.E., Xu, Z.(2004) Proc Natl Acad Sci U S A 101: 302-307
- PubMed: 14694194
- DOI: https://doi.org/10.1073/pnas.2036536100
- Primary Citation of Related Structures:
1UKF - PubMed Abstract:
AvrPphB is an avirulence (Avr) protein from the plant pathogen Pseudomonas syringae that can trigger a disease-resistance response in a number of host plants including Arabidopsis. AvrPphB belongs to a novel family of cysteine proteases with the charter member of this family being the Yersinia effector protein YopT. AvrPphB has a very stringent substrate specificity, catalyzing a single proteolytic cleavage in the Arabidopsis serine/threonine kinase PBS1. We have determined the crystal structure of AvrPphB by x-ray crystallography at 1.35-A resolution. The structure is composed of a central antiparallel beta-sheet, with alpha-helices packing on both sides of the sheet to form a two-lobe structure. The core of this structure resembles the papain-like cysteine proteases. The similarity includes the AvrPphB active site catalytic triad of Cys-98, His-212, and Asp-227 and the oxyanion hole residue Asn-93. Based on analogy with inhibitor complexes of the papain-like proteases, we propose a model for the substrate-binding mechanism of AvrPphB. A deep and positively charged pocket (S2) and a neighboring shallow surface (S3) likely bind to aspartic acid and glycine residues in the substrate located two (P2) and three (P3) residues N terminal to the cleavage site, respectively. Further implications about the specificity of plant pathogen recognition are also discussed.
Organizational Affiliation:
Department of Biological Chemistry, Medical School and Life Sciences Institute, University of Michigan, Ann Arbor, MI 48109, USA.