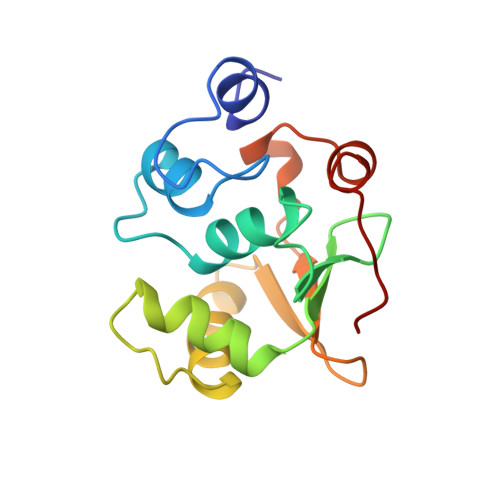
The Structural and Dynamic Basis of Ets-1 DNA Binding Autoinhibition
Lee, G.M., Donaldson, L.W., Pufall, M.A., Kang, H.-S., Pot, I., Graves, B.J., McIntosh, L.P.(2005) J Biol Chem 280: 7088-7099
- PubMed: 15591056
- DOI: https://doi.org/10.1074/jbc.M410722200
- Primary Citation of Related Structures:
1R36 - PubMed Abstract:
The transcription factor Ets-1 is regulated by the allosteric coupling of DNA binding with the unfolding of an alpha-helix (HI-1) within an autoinhibitory module. To understand the structural and dynamic basis for this autoinhibition, we have used NMR spectroscopy to characterize Ets-1DeltaN301, a partially inhibited fragment of Ets-1. The NMR-derived Ets-1DeltaN301 structure reveals that the autoinhibitory module is formed predominantly by the hydrophobic packing of helices from the N-terminal (HI-1, HI-2) and C-terminal (H4, H5) inhibitory sequences, along with H1 of the intervening DNA binding ETS domain. The intramolecular interactions made by HI-1 in Ets-1DeltaN301 are similar to the intermolecular contacts observed in the crystal structure of an Ets-1DeltaN300 dimer, confirming that the latter represents a domain-swapped species. (15)N relaxation studies demonstrate that the backbone of the N-terminal inhibitory sequence is mobile on the nanosecond-picosecond and millisecond-microsecond time scales. Furthermore, hydrogen exchange measurements reveal that amide protons in helices HI-1 and HI-2 exchange with water at rates only approximately 15- and approximately 75-fold slower, respectively, than predicted for an unfolded polypeptide. These findings indicate that inhibitory helices are only marginally stable even in the absence of DNA. The energetic coupling of DNA binding with the facile unfolding of the labile HI-1 provides a mechanism for modulating Ets-1 DNA binding activity via protein partnerships, post-translational modifications, or mutations. Ets-1 autoinhibition illustrates how conformational equilibria within structural domains can regulate macromolecular interactions.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of British Columbia, Vancouver, British Columbia V6T 1Z3, Canada.