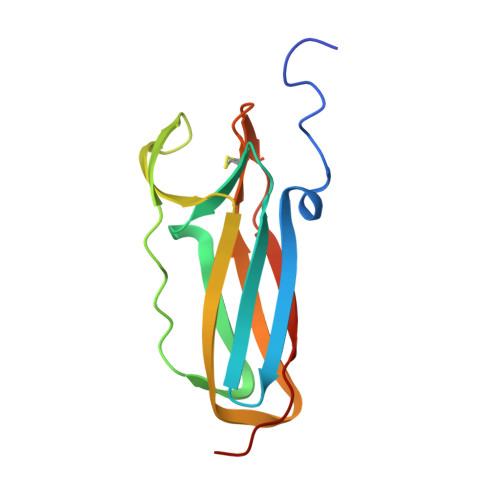
Thiol-disulfide exchange in an immunoglobulin-like fold: structure of the N-terminal domain of DsbD.
Goulding, C.W., Sawaya, M.R., Parseghian, A., Lim, V., Eisenberg, D., Missiakas, D.(2002) Biochemistry 41: 6920-6927
- PubMed: 12033924
- DOI: https://doi.org/10.1021/bi016038l
- Primary Citation of Related Structures:
1L6P - PubMed Abstract:
Escherichia coli DsbD transports electrons across the plasma membrane, a pathway that leads to the reduction of protein disulfide bonds. Three secreted thioredoxin-like factors, DsbC, DsbE, and DsbG, reduce protein disulfide bonds whereby an active site C-X-X-C motif is oxidized to generate a disulfide bond. DsbD catalyzes the reduction of the disulfide of DsbC, DsbE, and DsbG but not of the thioredoxin-like oxidant DsbA. The reduction of DsbC, DsbE, and DsbG occurs by transport of electrons from cytoplasmic thioredoxin to the C-terminal thioredoxin-like domain of DsbD (DsbD(C)). The N-terminal domain of DsbD, DsbD(N), acts as a versatile adaptor in electron transport and is capable of forming disulfides with oxidized DsbC, DsbE, or DsbG as well as with reduced DsbD(C). Isolated DsbD(N) is functional in electron transport in vitro. Crystallized DsbD(N) assumes an immunoglobulin-like fold that encompasses two active site cysteines, C103 and C109, forming a disulfide bond between beta-strands. The disulfide of DsbD(N) is shielded from the environment and capped by a phenylalanine (F70). A model is discussed whereby the immunoglobulin fold of DsbD(N) may provide for the discriminating interaction with thioredoxin-like factors, thereby triggering movement of the phenylalanine cap followed by disulfide rearrangement.
Organizational Affiliation:
Howard Hughes Medical Institute and Laboratory of Structural Biology and Molecular Medicine, UCLA-DOE, P.O. Box 951570, Los Angeles, California 90095-1570, USA.