Structure and Mechanism of Escherichia coli Aminodeoxychorismate Lyase
Jensen, P.Y., Parsons, J.F., Fisher, K.E., Pachikara, A.S., Tordova, M., Howard, A.J., Eisenstein, E., Ladner, J.E.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 4-AMINO-4-DEOXYCHORISMATE LYASE | 269 | Escherichia coli | Mutation(s): 0 Gene Names: PABC EC: 4 | 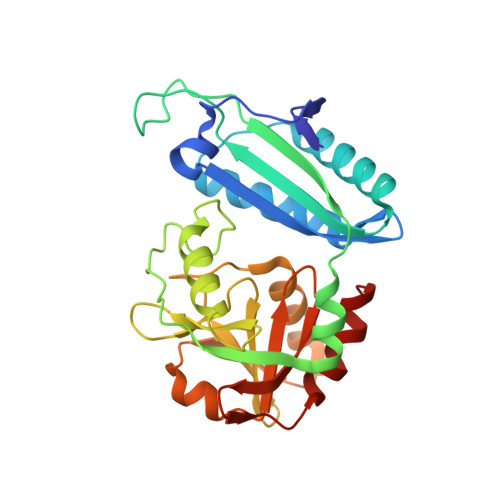 | |
UniProt | |||||
Find proteins for P28305 (Escherichia coli (strain K12)) Explore P28305 Go to UniProtKB: P28305 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P28305 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PLP Query on PLP | B [auth A] | PYRIDOXAL-5'-PHOSPHATE C8 H10 N O6 P NGVDGCNFYWLIFO-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 40.047 | α = 90 |
| b = 72.966 | β = 90 |
| c = 82.871 | γ = 90 |
| Software Name | Purpose |
|---|---|
| X-GEN | data scaling |
| X-GEN | data reduction |
| SHELX | model building |
| SHELXL-97 | refinement |
| SHELX | phasing |