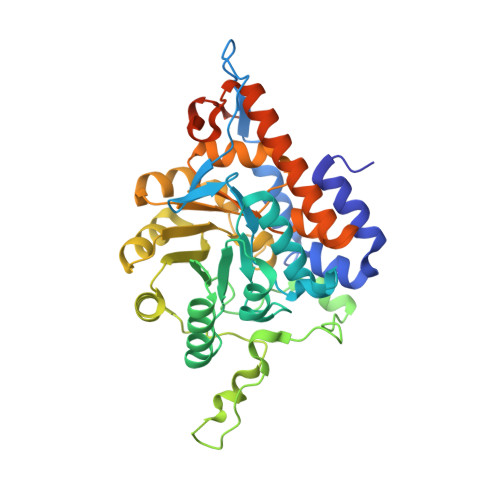
Structure of an active soluble mutant of the membrane-associated (S)-mandelate dehydrogenase.
Sukumar, N., Xu, Y., Gatti, D.L., Mitra, B., Mathews, F.S.(2001) Biochemistry 40: 9870-9878
- PubMed: 11502180
- DOI: https://doi.org/10.1021/bi010938k
- Primary Citation of Related Structures:
1HUV - PubMed Abstract:
The structure of an active mutant of (S)-mandelate dehydrogenase (MDH-GOX2) from Pseudomonas putida has been determined at 2.15 A resolution. The membrane-associated flavoenzyme (S)-mandelate dehydrogenase (MDH) catalyzes the oxidation of (S)-mandelate to give a flavin hydroquinone intermediate which is subsequently reoxidized by an organic oxidant residing in the membrane. The enzyme was rendered soluble by replacing its 39-residue membrane-binding peptide segment with a corresponding 20-residue segment from its soluble homologue, glycolate oxidase (GOX). Because of their amphipathic nature and peculiar solubilization properties, membrane proteins are notoriously difficult to crystallize, yet represent a large fraction of the proteins encoded by genomes currently being deciphered. Here we present the first report of such a structure in which an internal membrane-binding segment has been replaced, leading to successful crystallization of the fully active enzyme in the absence of detergents. This approach may have general application to other membrane-bound proteins. The overall fold of the molecule is that of a TIM barrel, and it forms a tight tetramer within the crystal lattice that has circular 4-fold symmetry. The structure of MDH-GOX2 reveals how this molecule can interact with a membrane, although it is limited by the absence of a membrane-binding segment. MDH-GOX2 and GOX adopt similar conformations, yet they retain features characteristic of membrane and globular proteins, respectively. MDH-GOX2 has a distinctly electropositive surface capable of interacting with the membrane, while the opposite surface is largely electronegative. GOX shows no such pattern. MDH appears to form a new class of monotopic integral membrane protein that interacts with the membrane through coplanar electrostatic binding surfaces and hydrophobic interactions, thus combining features of both the prostaglandin synthase/squaline-hopine cyclase and the C-2 coagulation factor domain classes of membrane proteins.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, Missouri 63110, USA.