A new mode of B12 binding and the direct participation of a potassium ion in enzyme catalysis: X-ray structure of diol dehydratase.
Shibata, N., Masuda, J., Tobimatsu, T., Toraya, T., Suto, K., Morimoto, Y., Yasuoka, N.(1999) Structure 7: 997-1008
- PubMed: 10467140
- DOI: https://doi.org/10.1016/s0969-2126(99)80126-9
- Primary Citation of Related Structures:
1DIO - PubMed Abstract:
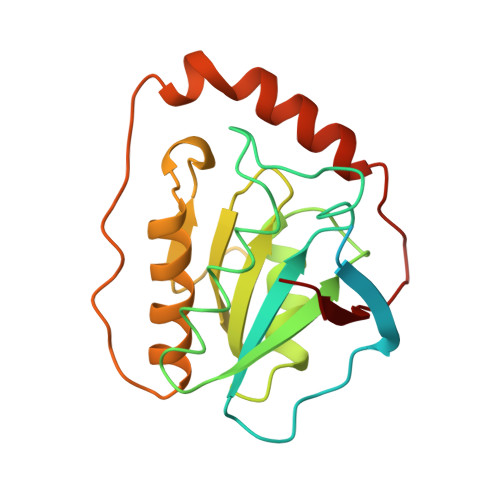
Diol dehydratase is an enzyme that catalyzes the adenosylcobalamin (coenzyme B12) dependent conversion of 1,2-diols to the corresponding aldehydes. The reaction initiated by homolytic cleavage of the cobalt-carbon bond of the coenzyme proceeds by a radical mechanism. The enzyme is an alpha2beta2gamma2 heterooligomer and has an absolute requirement for a potassium ion for catalytic activity. The crystal structure analysis of a diol dehydratase-cyanocobalamin complex was carried out in order to help understand the mechanism of action of this enzyme. The three-dimensional structure of diol dehydratase in complex with cyanocobalamin was determined at 2.2 A resolution. The enzyme exists as a dimer of heterotrimers (alphabetagamma)2. The cobalamin molecule is bound between the alpha and beta subunits in the 'base-on' mode, that is, 5,6-dimethylbenzimidazole of the nucleotide moiety coordinates to the cobalt atom in the lower axial position. The alpha subunit includes a (beta/alpha)8 barrel. The substrate, 1,2-propanediol, and an essential potassium ion are deeply buried inside the barrel. The two hydroxyl groups of the substrate coordinate directly to the potassium ion. This is the first crystallographic indication of the 'base-on' mode of cobalamin binding. An unusually long cobalt-base bond seems to favor homolytic cleavage of the cobalt-carbon bond and therefore to favor radical enzyme catalysis. Reactive radical intermediates can be protected from side reactions by spatial isolation inside the barrel. On the basis of unique direct interactions between the potassium ion and the two hydroxyl groups of the substrate, direct participation of a potassium ion in enzyme catalysis is strongly suggested.
Organizational Affiliation:
Department of Life Science, Himeji Institute of Technology, Hyogo, Japan.