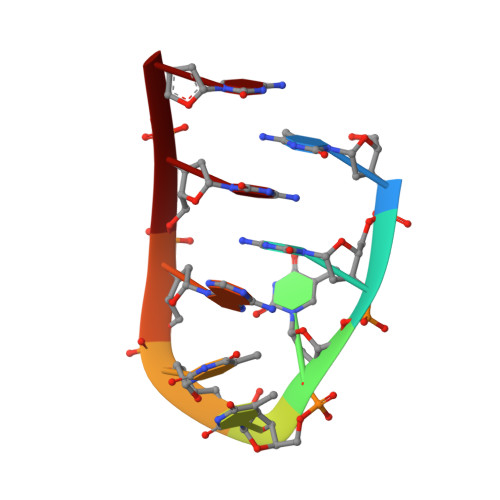
Solution structure and base pair opening kinetics of the i-motif dimer of d(5mCCTTTACC): a noncanonical structure with possible roles in chromosome stability.
Nonin, S., Phan, A.T., Leroy, J.L.(1997) Structure 5: 1231-1246
- PubMed: 9331414
- DOI: https://doi.org/10.1016/s0969-2126(97)00273-6
- Primary Citation of Related Structures:
1BAE - PubMed Abstract:
Repetitive cytosine-rich DNA sequences have been identified in telomeres and centromeres of eukaryotic chromosomes. These sequences play a role in maintaining chromosome stability during replication and may be involved in chromosome pairing during meiosis. The C-rich repeats can fold into an 'i-motif' structure, in which two parallel-stranded duplexes with hemiprotonated C.C+ pairs are intercalated. Previous NMR studies of naturally occurring repeats have produced poor NMR spectra. This led us to investigate oligonucleotides, based on natural sequences, to produce higher quality spectra and thus provide further information as to the structure and possible biological function of the i-motif.
Organizational Affiliation:
Groupe de Biophysique, de l'Ecole Polytechnique et de l'URA, CNRS, Palaiseau, France.