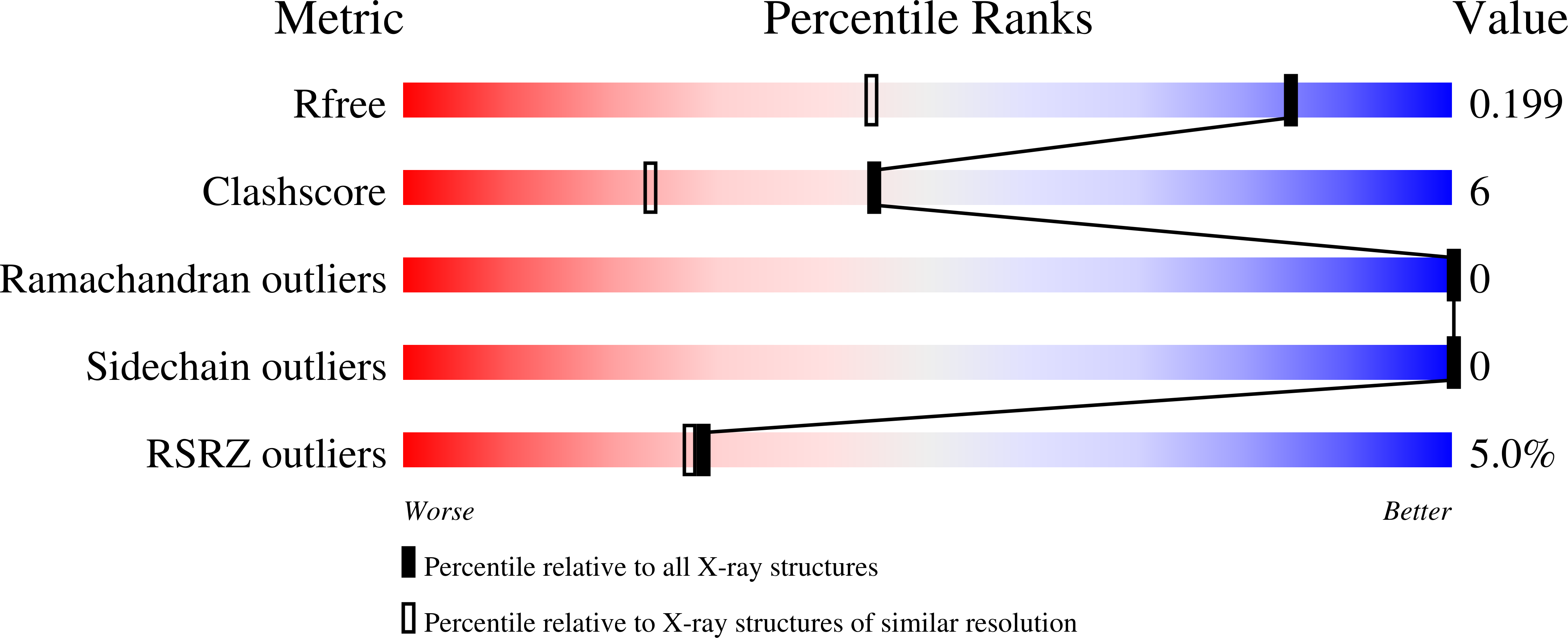
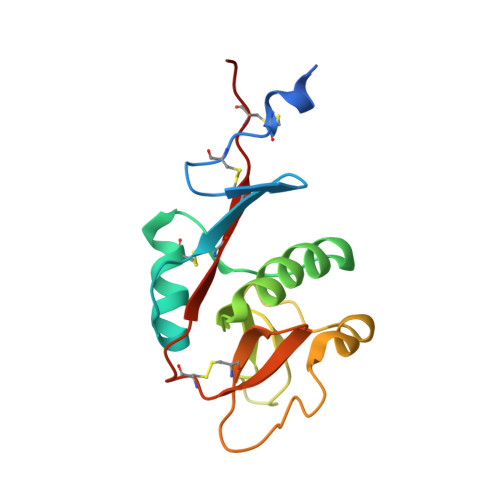
The structure of DC-SIGNR with a portion of its repeat domain lends insights to modeling of the receptor tetramer.
Snyder, G.A., Colonna, M., Sun, P.D.(2005) J Mol Biol 347: 979-989
- PubMed: 15784257
- DOI: https://doi.org/10.1016/j.jmb.2005.01.063
- Primary Citation of Related Structures:
1XPH - PubMed Abstract:
The dendritic cell-specific ICAM-3 non-integrin (DC-SIGN) and its close relative DC-SIGNR recognize various glycoproteins, both pathogenic and cellular, through the receptor lectin domain-mediated carbohydrate recognition. While the carbohydrate-recognition domains (CRD) exist as monomers and bind individual carbohydrates with low affinity and are permissive in nature, the full-length receptors form tetramers through their repeat domain and recognize specific ligands with high affinity. To understand the tetramer-based ligand binding avidity, we determined the crystal structure of DC-SIGNR with its last repeat region. Compared to the carbohydrate-bound CRD structure, the structure revealed conformational changes in the calcium and carbohydrate coordination loops of CRD, an additional disulfide bond between the N and the C termini of the CRD, and a helical conformation for the last repeat. On the basis of the current crystal structure and other published structures with sequence homology to the repeat domain, we generated a tetramer model for DC-SIGN/R using homology modeling and propose a ligand-recognition index to identify potential receptor ligands.
Organizational Affiliation:
Laboratory of Immunogenetics, Structural Immunology Section, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Twinbrook II, 12441 Parklawn Drive, Rockville, MD 20852, USA.