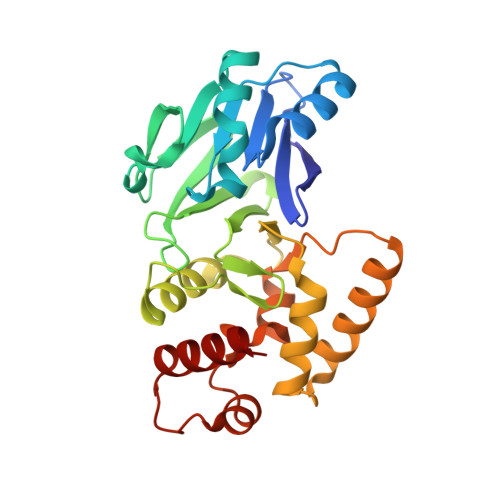
Structural studies on a mitochondrial glyoxalase II.
Marasinghe, G.P., Sander, I.M., Bennett, B., Periyannan, G., Yang, K.W., Makaroff, C.A., Crowder, M.W.(2005) J Biol Chem 280: 40668-40675
- PubMed: 16227621
- DOI: https://doi.org/10.1074/jbc.M509748200
- Primary Citation of Related Structures:
1XM8 - PubMed Abstract:
Glyoxalase 2 is a beta-lactamase fold-containing enzyme that appears to be involved with cellular chemical detoxification. Although the cytoplasmic isozyme has been characterized from several organisms, essentially nothing is known about the mitochondrial proteins. As a first step in understanding the structure and function of mitochondrial glyoxalase 2 enzymes, a mitochondrial isozyme (GLX2-5) from Arabidopsis thaliana was cloned, overexpressed, purified, and characterized using metal analyses, EPR and (1)H NMR spectroscopies, and x-ray crystallography. The recombinant enzyme was shown to bind 1.04 +/- 0.15 eq of iron and 1.31 +/- 0.05 eq of Zn(II) and to exhibit k(cat) and K(m) values of 129 +/- 10 s(-1) and 391 +/- 48 microm, respectively, when using S-d-lactoylglutathione as the substrate. EPR spectra revealed that recombinant GLX2-5 contains multiple metal centers, including a predominant Fe(III)Z-n(II) center and an anti-ferromagnetically coupled Fe(III)Fe(II) center. Unlike cytosolic glyoxalase 2 from A. thaliana, GLX2-5 does not appear to specifically bind manganese. (1)H NMR spectra revealed the presence of at least eight paramagnetically shifted resonances that arise from protons in close proximity to a Fe(III)Fe(II) center. Five of these resonances arose from solvent-exchangeable protons, and four of these have been assigned to NH protons on metal-bound histidines. A 1.74-A resolution crystal structure of the enzyme revealed that although GLX2-5 shares a number of structural features with human GLX2, several important differences exist. These data demonstrate that mitochondrial glyoxalase 2 can accommodate a number of different metal centers and that the predominant metal center is Fe(III)Zn(II).
Organizational Affiliation:
Department of Chemistry and Biochemistry, Miami University, Oxford, Ohio 45056, USA.