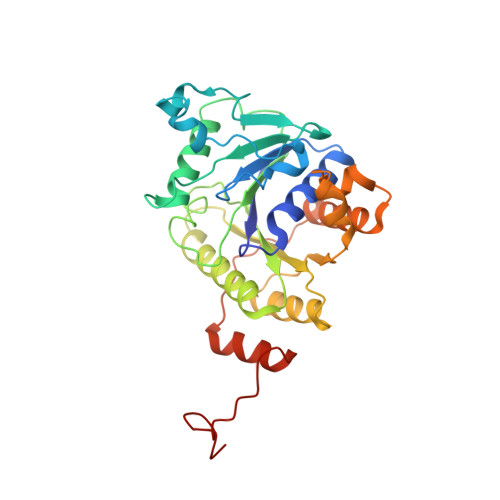
Structural basis for the changes in redox potential in the nitrogenase Phe135Trp Fe protein with MgADP Bound
Jeong, M.S., Jang, S.B.(2004) Mol Cell 18: 374-382
- PubMed: 15650336
- Primary Citation of Related Structures:
1XCP - PubMed Abstract:
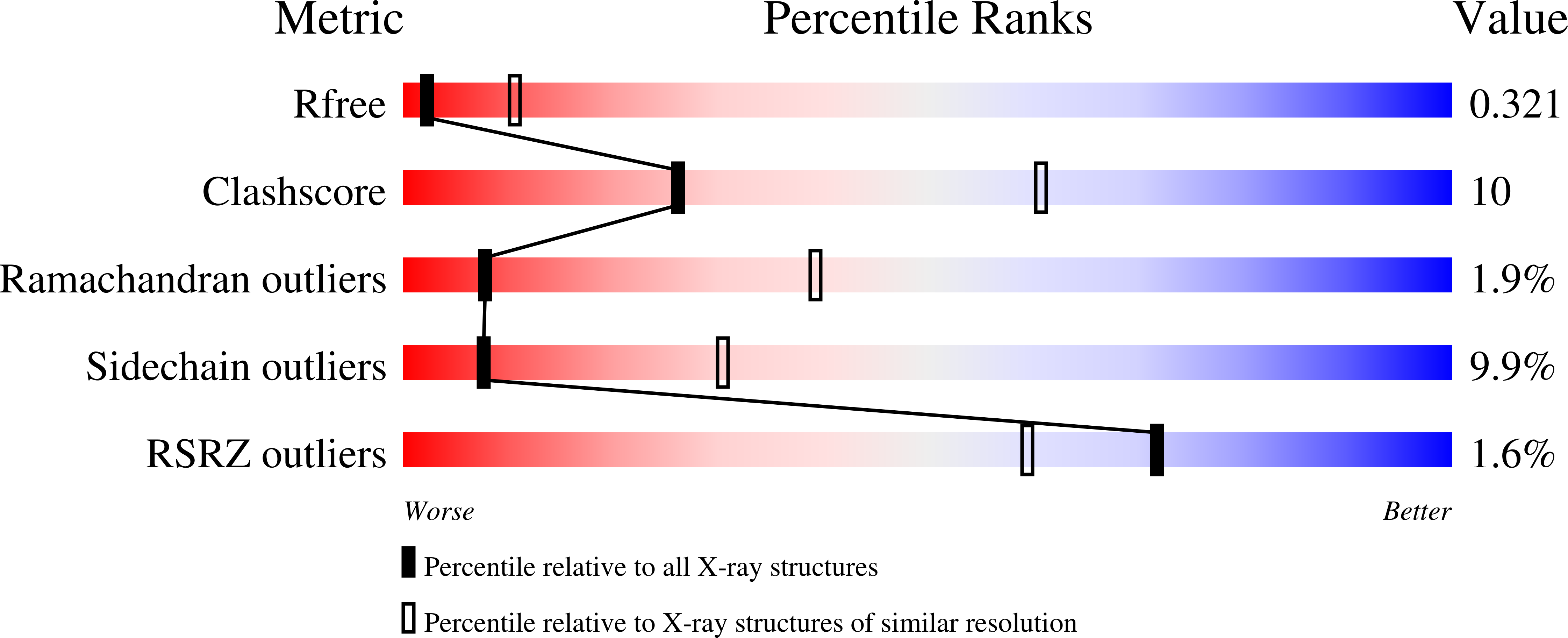
The crystal structure of the Azotobacter vinelandii nitrogenase Fe protein with phenylalanine at position 135 substituted by tryptophan has been determined in MgADP-bound form by X-ray diffraction methods. Amino acid substitution studies have suggested that the phenylalanine at position 135 located near the [4Fe-4S] cluster contributes to both the midpoint potential and nucleotide-induced changes of the [4Fe-4S] cluster. Substitution of tryptophan for phenylalanine at position 135 resulted in a significant positive shift in the midpoint potential in both the isolated and nucleotide-bound states. The factors thought to control the midpoint potential of the [FeS] cluster include solvent accessibility, dipolar environment, and structural strain. The structure derived in the present work provides an explanation for the more positive midpoint potential observed in the nucleotide-bound state, and suggests important insights into the contributions of the nucleotide interaction to the conformational states that are the keys to nitrogenase catalysis. The presence of MgADP in Phe135Trp Fe protein reveals the mechanism of the long-range communication from the nucleotide-binding site that controls its affinity for the MoFe protein component.
Organizational Affiliation:
Korea Nanobiotechnology Center, Pusan National University, Busan 609-735, Korea.