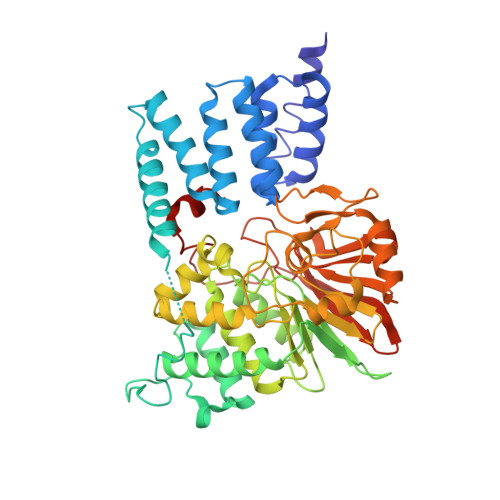
Molecular Basis for Tpr Domain-Mediated Regulation of Protein Phosphatase 5
Yang, J., Roe, S.M., Cliff, M.J., Williams, M.A., Ladbury, J.E., Cohen, P.T., Barford, D.(2005) EMBO J 24: 1
- PubMed: 15577939
- DOI: https://doi.org/10.1038/sj.emboj.7600496
- Primary Citation of Related Structures:
1WAO - PubMed Abstract:
Protein phosphatase 5 (Ppp5) is a serine/threonine protein phosphatase comprising a regulatory tetratricopeptide repeat (TPR) domain N-terminal to its phosphatase domain. Ppp5 functions in signalling pathways that control cellular responses to stress, glucocorticoids and DNA damage. Its phosphatase activity is suppressed by an autoinhibited conformation maintained by the TPR domain and a C-terminal subdomain. By interacting with the TPR domain, heat shock protein 90 (Hsp90) and fatty acids including arachidonic acid stimulate phosphatase activity. Here, we describe the structure of the autoinhibited state of Ppp5, revealing mechanisms of TPR-mediated phosphatase inhibition and Hsp90- and arachidonic acid-induced stimulation of phosphatase activity. The TPR domain engages with the catalytic channel of the phosphatase domain, restricting access to the catalytic site. This autoinhibited conformation of Ppp5 is stabilised by the C-terminal alphaJ helix that contacts a region of the Hsp90-binding groove on the TPR domain. Hsp90 activates Ppp5 by disrupting TPR-phosphatase domain interactions, permitting substrate access to the constitutively active phosphatase domain, whereas arachidonic acid prompts an alternate conformation of the TPR domain, destabilising the TPR-phosphatase domain interface.
Organizational Affiliation:
Section of Structural Biology, The Institute of Cancer Research, Chester Beatty Laboratories, London, UK.