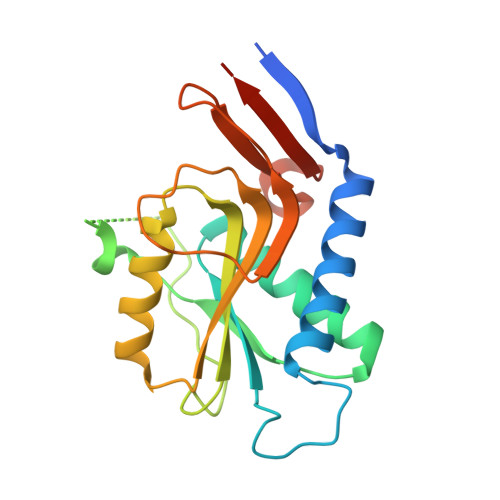
Structure of Pyrr (Rv1379) from Mycobacterium Tuberculosis: A Persistence Gene and Protein Drug Target
Kantardjieff, K.A., Vasquez, C., Castro, P., Warfel, N.N., Rho, B.-S., Lekin, T., Kim, C.-Y., Segelke, B.W., Terwilliger, T., Rupp, B.(2005) Acta Crystallogr D Biol Crystallogr 61: 355
- PubMed: 15805589
- DOI: https://doi.org/10.1107/S090744490403389X
- Primary Citation of Related Structures:
1W30 - PubMed Abstract:
The Mycobacterium tuberculosis pyrR gene (Rv1379) encodes a protein that regulates the expression of pyrimidine-nucleotide biosynthesis (pyr) genes in a UMP-dependent manner. Because pyrimidine biosynthesis is an essential step in the progression of TB, the gene product pyrR is an attractive antitubercular drug target. The 1.9 A native structure of Mtb pyrR determined by the TB Structural Genomics Consortium facilities in trigonal space group P3(1)21 is reported, with unit-cell parameters a = 66.64, c = 154.72 A at 120 K and two molecules in the asymmetric unit. The three-dimensional structure and residual uracil phosphoribosyltransferase activity point to a common PRTase ancestor for pyrR. However, while PRPP- and UMP-binding sites have been retained in Mtb pyrR, a distinct dimer interaction among subunits creates a deep positively charged cleft capable of binding pyr mRNA. In silico screening of pyrimidine-nucleoside analogs has revealed a number of potential lead compounds that, if bound to Mtb pyrR, could facilitate transcriptional attenuation, particularly cyclopentenyl nucleosides.
Organizational Affiliation:
Department of Chemistry and Biochemistry and W. M. Keck Foundation Center for Molecular Structure, California State University Fullerton, Fullerton, CA 92834, USA.