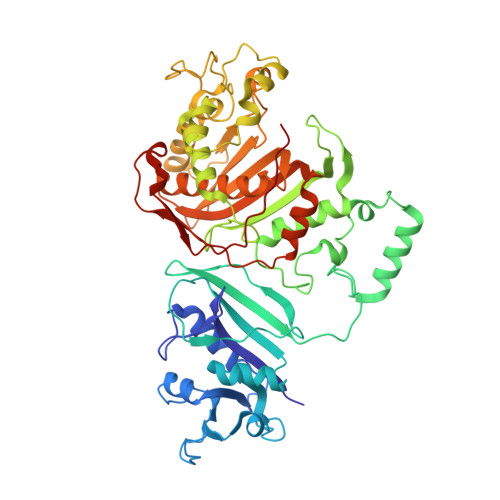
Two crystal structures of dihydrofolate reductase-thymidylate synthase from Cryptosporidium hominis reveal protein-ligand interactions including a structural basis for observed antifolate resistance.
Anderson, A.C.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 258-262
- PubMed: 16511011
- DOI: https://doi.org/10.1107/S1744309105002435
- Primary Citation of Related Structures:
1SEJ - PubMed Abstract:
Cryptosporidium hominis is a protozoan parasite that causes acute gastrointestinal illness. There are no effective therapies for cryptosporidiosis, highlighting the need for new drug-lead discovery. An analysis of the protein-ligand interactions in two crystal structures of dihydrofolate reductase-thymidylate synthase (DHFR-TS) from C. hominis, determined at 2.8 and 2.87 A resolution, reveals that the interactions of residues Ile29, Thr58 and Cys113 in the active site of C. hominis DHFR provide a possible structural basis for the observed antifolate resistance. A comparison with the structure of human DHFR reveals active-site differences that may be exploited for the design of species-selective inhibitors.
Organizational Affiliation:
Department of Chemistry, Dartmouth College, Hanover, NH 03755, USA. aca@dartmouth.edu