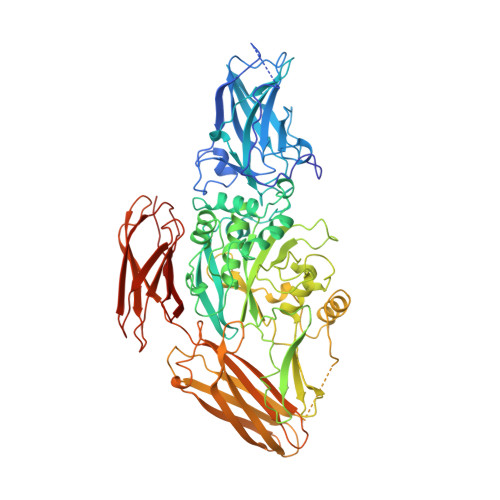
Identification of the calcium binding site and a novel ytterbium site in blood coagulation factor XIII by x-ray crystallography.
Fox, B.A., Yee, V.C., Pedersen, L.C., Le Trong, I., Bishop, P.D., Stenkamp, R.E., Teller, D.C.(1999) J Biol Chem 274: 4917-4923
- PubMed: 9988734
- DOI: https://doi.org/10.1074/jbc.274.8.4917
- Primary Citation of Related Structures:
1GGU, 1GGY, 1QRK - PubMed Abstract:
The presence or absence of calcium determines the activation, activity, oligomerization, and stability of blood coagulation factor XIII. To explore these observed effects, we have determined the x-ray crystal structure of recombinant factor XIII A2 in the presence of calcium, strontium, and ytterbium. The main calcium binding site within each monomer involves the main chain oxygen atom of Ala-457, and also the side chains from residues Asn-436, Asp-438, Glu-485, and Glu-490. Calcium and strontium bind in the same location, while ytterbium binds several angstroms removed. A novel ytterbium binding site is also found at the dimer two-fold axis, near residues Asp-270 and Glu-272, and this site may be related to the reported inhibition by lanthanide metals (Achyuthan, K. E., Mary, A., and Greenberg, C. S. (1989) Biochem. J. 257, 331-338). The overall structure of ion-bound factor XIII is very similar to the previously determined crystal structures of factor XIII zymogen, likely due to the constraints of this monoclinic crystal form. We have merged the three independent sets of water molecules in the structures to determine which water molecules are conserved and possibly structurally significant.
Organizational Affiliation:
Department of Biochemistry, University of Washington, Seattle, Washington 98195, USA.