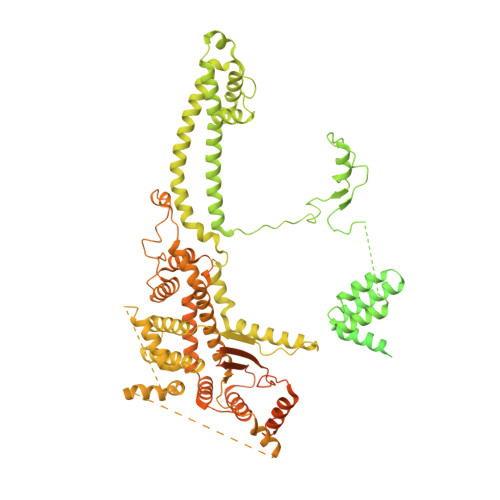
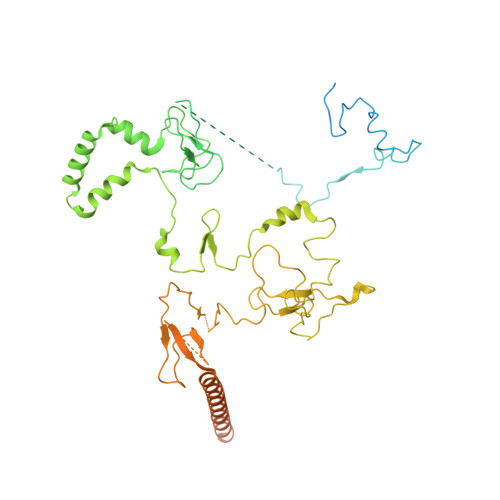
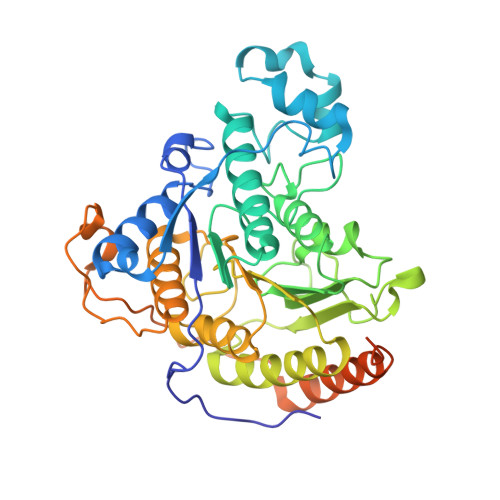
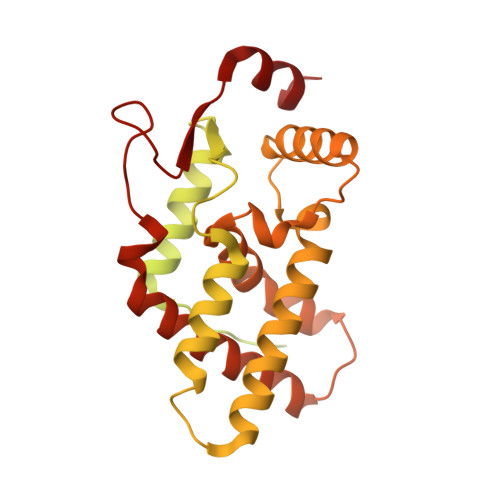
Structures and dynamics of Rpd3S complex bound to nucleosome.
Wang, C., Chu, C., Guo, Z., Zhan, X.(2024) Sci Adv 10: eadk7678-eadk7678
- PubMed: 38598631
- DOI: https://doi.org/10.1126/sciadv.adk7678
- Primary Citation of Related Structures:
8W9C, 8W9D, 8W9E, 8W9F - PubMed Abstract:
The Rpd3S complex plays a pivotal role in facilitating local histone deacetylation in the transcribed regions to suppress intragenic transcription initiation. Here, we present the cryo-electron microscopy structures of the budding yeast Rpd3S complex in both its apo and three nucleosome-bound states at atomic resolutions, revealing the exquisite architecture of Rpd3S to well accommodate a mononucleosome without linker DNA. The Rpd3S core, containing a Sin3 Lobe and two NB modules, is a rigid complex and provides three positive-charged anchors (Sin3_HCR and two Rco1_NIDs) to connect nucleosomal DNA. In three nucleosome-bound states, the Rpd3S core exhibits three distinct orientations relative to the nucleosome, assisting the sector-shaped deacetylase Rpd3 to locate above the SHL5-6, SHL0-1, or SHL2-3, respectively. Our work provides a structural framework that reveals a dynamic working model for the Rpd3S complex to engage diverse deacetylation sites.
Organizational Affiliation:
Westlake Laboratory of Life Sciences and Biomedicine, 18 Shilongshan Road, Hangzhou 310024, Zhejiang Province, China.