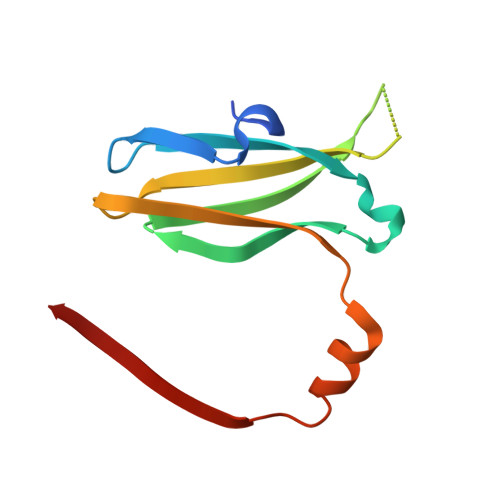
Mimicking a p53-MDM2 interaction based on a stable immunoglobulin-like domain scaffold.
Jimenez-Sandoval, P., Madrigal-Carrillo, E.A., Santamaria-Suarez, H.A., Maturana, D., Renteria-Gonzalez, I., Benitez-Cardoza, C.G., Torres-Larios, A., Brieba, L.G.(2018) Proteins 86: 802-812
- PubMed: 29696695
- DOI: https://doi.org/10.1002/prot.25519
- Primary Citation of Related Structures:
5SWK - PubMed Abstract:
Antibodies recognize protein targets with great affinity and specificity. However, posttranslational modifications and the presence of intrinsic disulfide-bonds pose difficulties for their industrial use. The immunoglobulin fold is one of the most ubiquitous folds in nature and it is found in many proteins besides antibodies. An example of a protein family with an immunoglobulin-like fold is the Cysteine Protease Inhibitors (ICP) family I42 of the MEROPs database for protease and protease inhibitors. Members of this protein family are thermostable and do not present internal disulfide bonds. Crystal structures of several ICPs indicate that they resemble the Ig-like domain of the human T cell co-receptor CD8α As ICPs present 2 flexible recognition loops that vary accordingly to their targeted protease, we hypothesize that members of this protein family would be ideal to design peptide aptamers that mimic protein-protein interactions. Herein, we use an ICP variant from Entamoeba histolytica (EhICP1) to mimic the interaction between p53 and MDM2. We found that a 13 amino-acid peptide derived from p53 can be introduced in 2 variable loops (DE, FG) but not the third (BC). Chimeric EhICP1-p53 form a stable complex with MDM2 at a micromolar range. Crystal structure of the EhICP1-p53(FG)-loop variant in complex with MDM2 reveals a swapping subdomain between 2 chimeric molecules, however, the p53 peptide interacts with MDM2 as in previous crystal structures. The structural details of the EhICP1-p53(FG) interaction with MDM2 resemble the interaction between an antibody and MDM2.
Organizational Affiliation:
Laboratorio Nacional de Genómica para la Biodiversidad, Centro de Investigación y de Estudios Avanzados del IPN, Apartado Postal 629, Irapuato, Guanajuato, CP 36821, México.