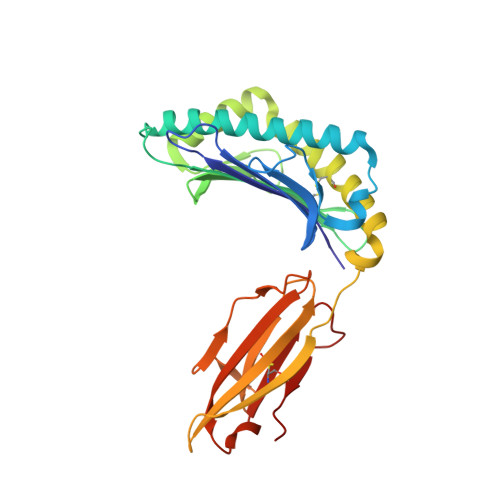
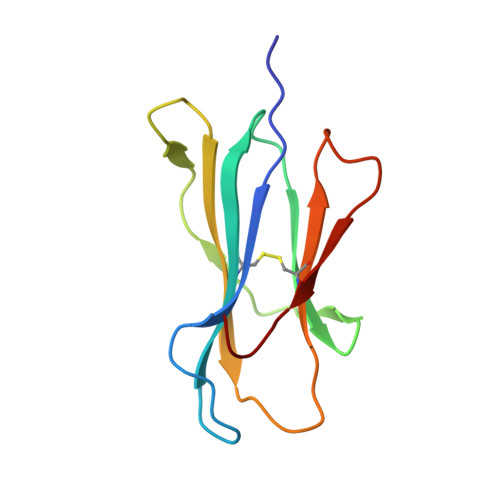
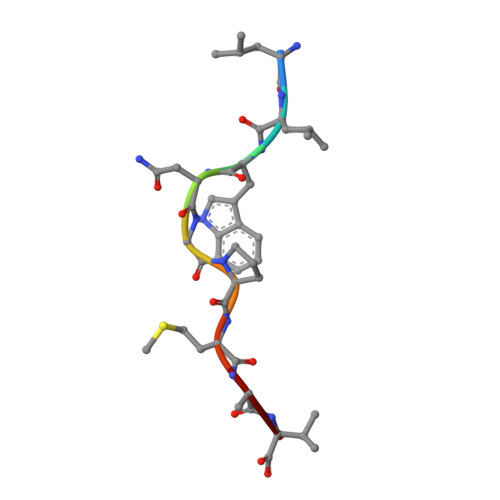
T cell receptor alpha variable 12-2 bias in the immunodominant response to Yellow fever virus.
Bovay, A., Zoete, V., Dolton, G., Bulek, A.M., Cole, D.K., Rizkallah, P.J., Fuller, A., Beck, K., Michielin, O., Speiser, D.E., Sewell, A.K., Fuertes Marraco, S.A.(2018) Eur J Immunol 48: 258-272
- PubMed: 28975614
- DOI: https://doi.org/10.1002/eji.201747082
- Primary Citation of Related Structures:
5N6B - PubMed Abstract:
The repertoire of human αβ T-cell receptors (TCRs) is generated via somatic recombination of germline gene segments. Despite this enormous variation, certain epitopes can be immunodominant, associated with high frequencies of antigen-specific T cells and/or exhibit bias toward a TCR gene segment. Here, we studied the TCR repertoire of the HLA-A*0201-restricted epitope LLWNGPMAV (hereafter, A2/LLW) from Yellow Fever virus, which generates an immunodominant CD8 + T cell response to the highly effective YF-17D vaccine. We discover that these A2/LLW-specific CD8 + T cells are highly biased for the TCR α chain TRAV12-2. This bias is already present in A2/LLW-specific naïve T cells before vaccination with YF-17D. Using CD8 + T cell clones, we show that TRAV12-2 does not confer a functional advantage on a per cell basis. Molecular modeling indicated that the germline-encoded complementarity determining region (CDR) 1α loop of TRAV12-2 critically contributes to A2/LLW binding, in contrast to the conventional dominant dependence on somatically rearranged CDR3 loops. This germline component of antigen recognition may explain the unusually high precursor frequency, prevalence and immunodominance of T-cell responses specific for the A2/LLW epitope.
Organizational Affiliation:
Department of Oncology, Lausanne University Hospital (CHUV), Epalinges, Switzerland.