Induced Structural Disorder as a Molecular Mechanism for Enzyme Dysfunction in Phosphoglucomutase 1 Deficiency.
Stiers, K.M., Kain, B.N., Graham, A.C., Beamer, L.J.(2016) J Mol Biol 428: 1493-1505
- PubMed: 26972339
- DOI: https://doi.org/10.1016/j.jmb.2016.02.032
- Primary Citation of Related Structures:
5EPC, 5F9C, 5HSH - PubMed Abstract:
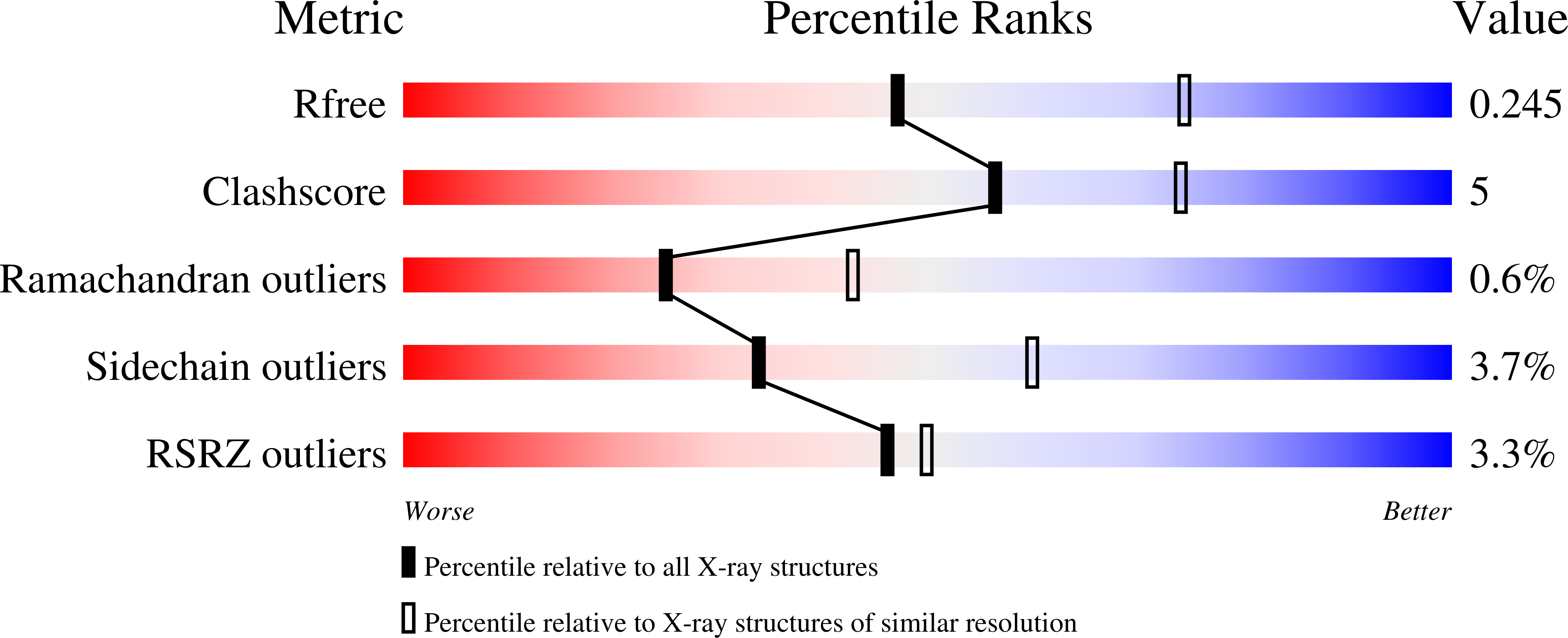
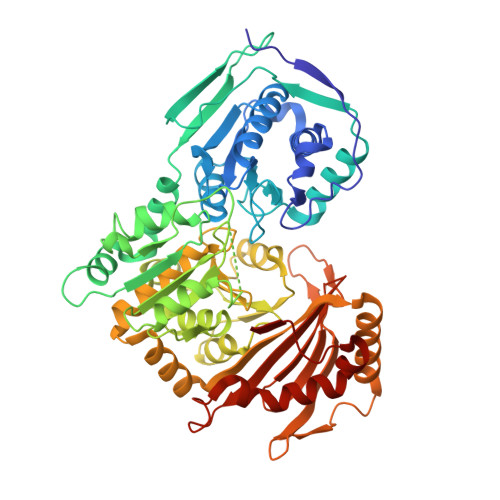
Human phosphoglucomutase 1 (PGM1) plays a central role in cellular glucose homeostasis, mediating the switch between glycolysis and gluconeogenesis through the conversion of glucose 1-phosphate and glucose 6-phosphate. Recent clinical studies have identified mutations in this enzyme as the cause of PGM1 deficiency, an inborn error of metabolism classified as both a glycogen storage disease and a congenital disorder of glycosylation. Reported here are the first crystal structures of two disease-related missense variants of PGM1, along with the structure of the wild-type enzyme. Two independent glycine-to-arginine substitutions (G121R and G291R), both affecting key active site loops of PGM1, are found to induce regions of structural disorder, as evidenced by a nearly complete loss of electron density for as many as 23 aa. The disordered regions are not contiguous in sequence to the site of mutation, and even cross domain boundaries. Other structural rearrangements include changes in the conformations of loops and side chains, some of which occur nearly 20 Å away from the site of mutation. The induced structural disorder is correlated with increased sensitivity to proteolysis and lower-resolution diffraction, particularly for the G291R variant. Examination of the multi-domain effects of these G➔R mutations establishes a correlation between interdomain interfaces of the enzyme and missense variants of PGM1 associated with disease. These crystal structures provide the first insights into the structural basis of enzyme dysfunction in PGM1 deficiency and highlight a growing role for biophysical characterization of proteins in the field of precision medicine.
Organizational Affiliation:
Biochemistry Department, University of Missouri, 117 Schweitzer Hall, Columbia, MO 65211, USA.