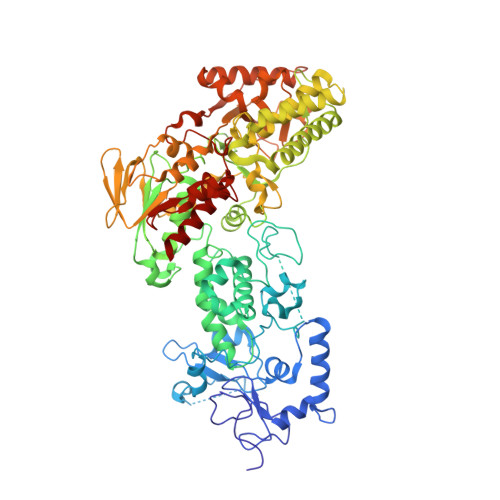
LSD2/KDM1B and its cofactor NPAC/GLYR1 endow a structural and molecular model for regulation of H3K4 demethylation
Fang, R., Chen, F., Dong, Z., Hu, D., Barbera, A.J., Clark, E.A., Fang, J., Yang, Y., Mei, P., Rutenberg, M., Li, Z., Zhang, Y., Xu, Y., Yang, H., Wang, P., Simon, M.D., Zhou, Q., Li, J., Marynick, M.P., Li, X., Lu, H., Kaiser, U.B., Kingston, R.E., Xu, Y., Shi, Y.G.(2013) Mol Cell 49: 558-570
- PubMed: 23260659
- DOI: https://doi.org/10.1016/j.molcel.2012.11.019
- Primary Citation of Related Structures:
4GU1, 4GUR, 4GUS, 4GUT, 4GUU - PubMed Abstract:
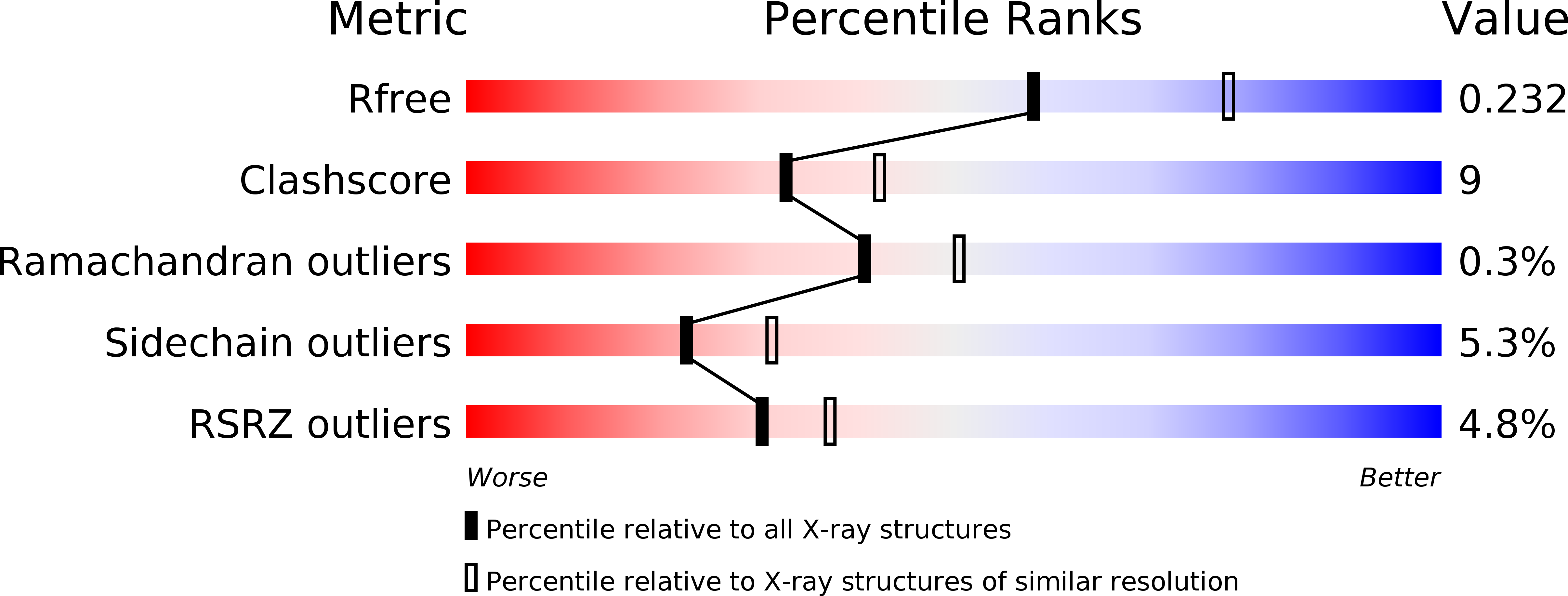
Dynamic regulation of histone methylation represents a fundamental epigenetic mechanism underlying eukaryotic gene regulation, yet little is known about how the catalytic activities of histone demethylases are regulated. Here, we identify and characterize NPAC/GLYR1 as an LSD2/KDM1b-specific cofactor that stimulates H3K4me1 and H3K4me2 demethylation. We determine the crystal structures of LSD2 alone and LSD2 in complex with the NPAC linker region in the absence or presence of histone H3 peptide, at resolutions of 2.9, 2.0, and 2.25 Å, respectively. These crystal structures and further biochemical characterization define a dodecapeptide of NPAC (residues 214-225) as the minimal functional unit for its cofactor activity and provide structural determinants and a molecular mechanism underlying the intrinsic cofactor activity of NPAC in stimulating LSD2-catalyzed H3K4 demethylation. Thus, these findings establish a model for how a cofactor directly regulates histone demethylation and will have a significant impact on our understanding of catalytic-activity-based epigenetic regulation.
Organizational Affiliation:
Division of Endocrinology, Diabetes, and Hypertension, Department of Medicine and Department of Biological Chemistry & Molecular Pharmacology, Brigham and Women's Hospital and Harvard Medical School, 221 Longwood Avenue, Boston, MA 02115, USA.