A single ligand is sufficient to activate EGFR dimers.
Liu, P., Cleveland, T.E., Bouyain, S., Byrne, P.O., Longo, P.A., Leahy, D.J.(2012) Proc Natl Acad Sci U S A 109: 10861-10866
- PubMed: 22699492
- DOI: https://doi.org/10.1073/pnas.1201114109
- Primary Citation of Related Structures:
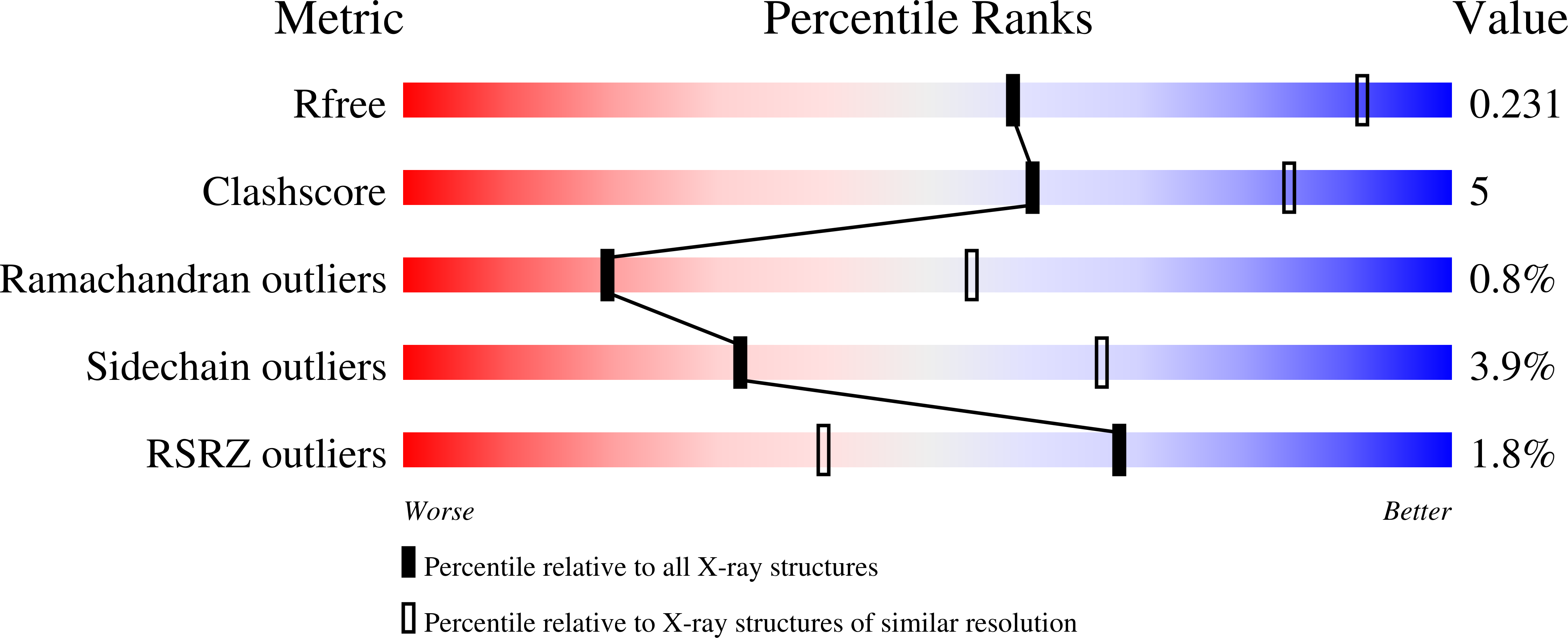
3U7U - PubMed Abstract:
Crystal structures of human epidermal growth factor receptor (EGFR) with bound ligand revealed symmetric, doubly ligated receptor dimers thought to represent physiologically active states. Such complexes fail to rationalize negative cooperativity of epidermal growth factor (EGF) binding to EGFR and the behavior of the ligandless EGFR homolog ErbB2/HER2, however. We report cell-based assays that provide evidence for active, singly ligated dimers of human EGFR and its homolog, ErbB4/HER4. We also report crystal structures of the ErbB4/HER4 extracellular region complexed with its ligand Neuregulin-1β that resolve two types of ErbB dimer when compared to EGFR:Ligand complexes. One type resembles the recently reported asymmetric dimer of Drosophila EGFR with a single high-affinity ligand bound and provides a model for singly ligated human ErbB dimers. These results unify models of vertebrate and invertebrate EGFR/ErbB signaling, imply that the tethered conformation of unliganded ErbBs evolved to prevent crosstalk among ErbBs, and establish a molecular basis for both negative cooperativity of ligand binding to vertebrate ErbBs and the absence of active ErbB2/HER2 homodimers in normal conditions.
Organizational Affiliation:
Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, 725 N. Wolfe Street, Baltimore, MD 21205, USA.