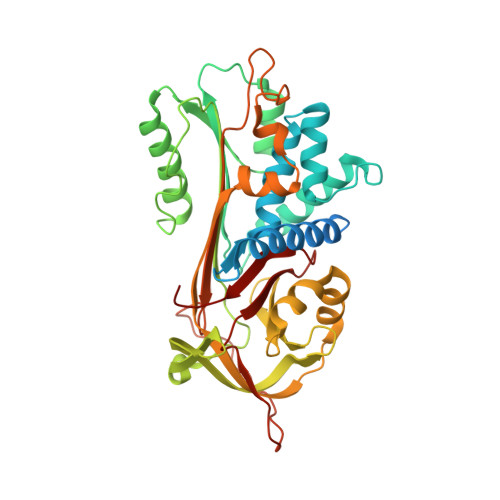
Therapeutic target-site variability in [alpha]1-antitrypsin characterized at high resolution
Patschull, A.O., Segu, L., Nyon, M.P., Lomas, D.A., Nobeli, I., Barrett, T.E., Gooptu, B.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1492-1497
- PubMed: 22139150
- DOI: https://doi.org/10.1107/S1744309111040267
- Primary Citation of Related Structures:
3NE4 - PubMed Abstract:
The intrinsic propensity of α(1)-antitrypsin to undergo conformational transitions from its metastable native state to hyperstable forms provides a motive force for its antiprotease function. However, aberrant conformational change can also occur via an intermolecular linkage that results in polymerization. This has both loss-of-function and gain-of-function effects that lead to deficiency of the protein in human circulation, emphysema and hepatic cirrhosis. One of the most promising therapeutic strategies being developed to treat this disease targets small molecules to an allosteric site in the α(1)-antitrypsin molecule. Partial filling of this site impedes polymerization without abolishing function. Drug development can be improved by optimizing data on the structure and dynamics of this site. A new 1.8 Å resolution structure of α(1)-antitrypsin demonstrates structural variability within this site, with associated fluctuations in its upper and lower entrance grooves and ligand-binding characteristics around the innermost stable enclosed hydrophobic recess. These data will allow a broader selection of chemotypes and derivatives to be tested in silico and in vitro when screening and developing compounds to modulate conformational change to block the pathological mechanism while preserving function.
Organizational Affiliation:
Institute of Structural and Molecular Biology, Crystallography, Department of Biological Sciences, Birkbeck College, London, England.