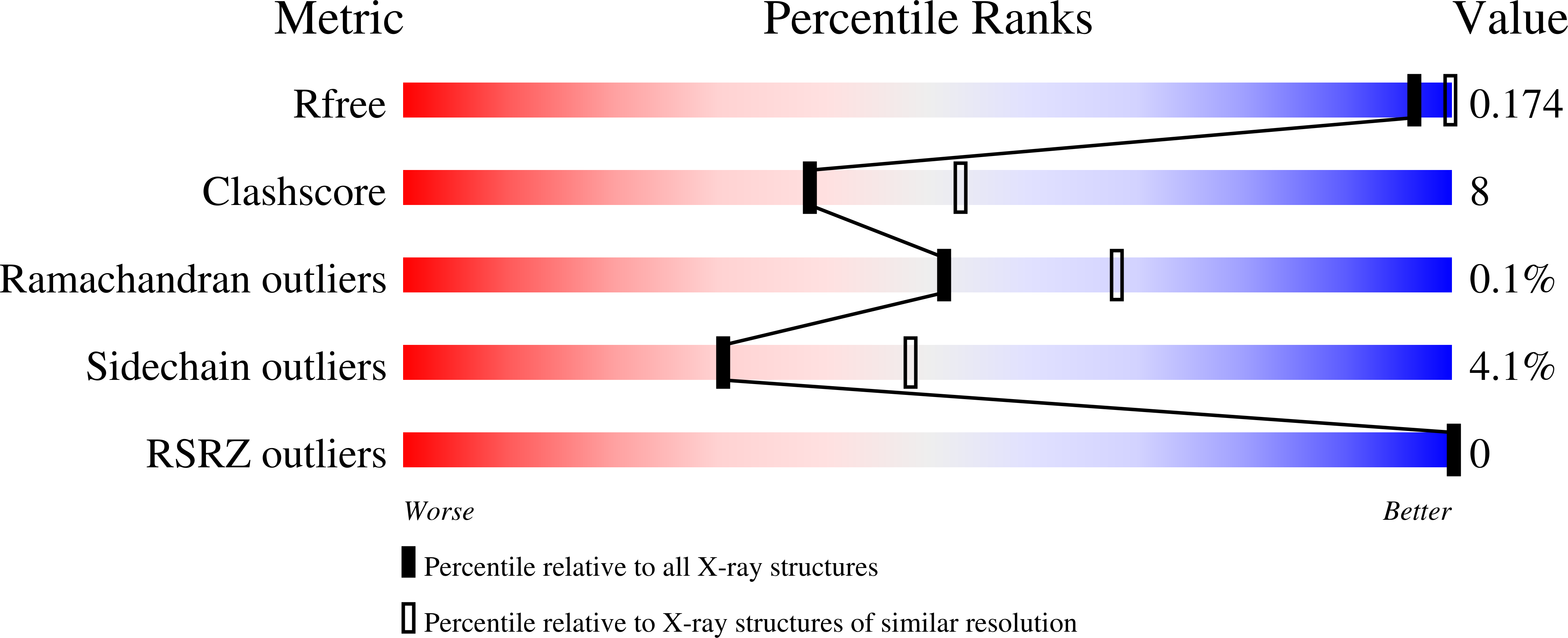
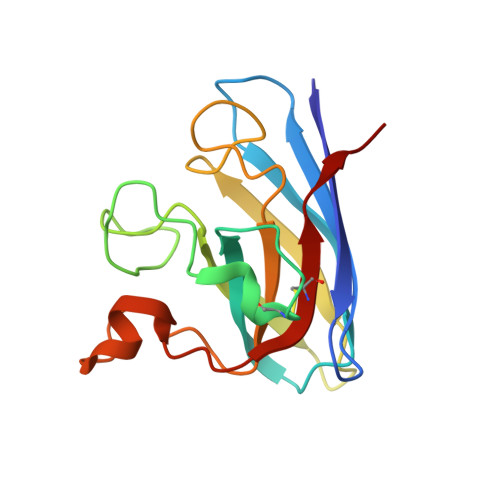
Structures of mouse SOD1 and human/mouse SOD1 chimeras.
Seetharaman, S.V., Taylor, A.B., Holloway, S., Hart, P.J.(2010) Arch Biochem Biophys 503: 183-190
- PubMed: 20727846
- DOI: https://doi.org/10.1016/j.abb.2010.08.014
- Primary Citation of Related Structures:
3GTT, 3GTV, 3LTV - PubMed Abstract:
Mutations in human copper-zinc superoxide dismutase (SOD1) cause an inherited form of amyotrophic lateral sclerosis (ALS). Inclusions enriched in pathogenic SOD1 accumulate in the spinal cords of transgenic mice expressing these proteins, but endogenous mouse SOD1 is not found as a component of these aggregates. In the accompanying paper, Karch and colleagues analyze aggregation propensities of human/mouse SOD1 chimeras in cell culture and identify two sequence elements in the human enzyme that seem to enhance its aggregation relative to the mouse enzyme. Here, we report the first structure of mouse SOD1 along with those of SOD1 chimeras in which residues 1-80 come from human SOD1 and residues 81-153 come from mouse SOD1 and vice versa. Taken together, the structural and cell-based data suggest a model in which residues Q42 and Q123 in mouse SOD1 modulate non-native SOD1-SOD1 intermolecular interactions at edge strands in the SOD1 Greek key β-barrel.
Organizational Affiliation:
Department of Biochemistry, The University of Texas Health Science Center, San Antonio, 78229, USA.