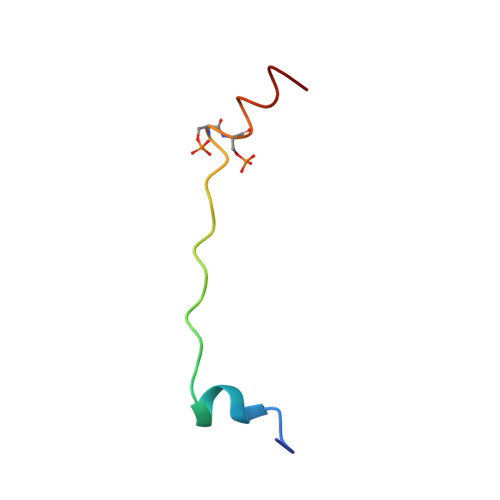
Phosphorylation-dependent conformational transition of the cardiac specific N-extension of troponin I in cardiac troponin
Howarth, J.W., Meller, J., Solaro, R.J., Trewhella, J., Rosevear, P.R.(2007) J Mol Biol 373: 706-722
- PubMed: 17854829
- DOI: https://doi.org/10.1016/j.jmb.2007.08.035
- Primary Citation of Related Structures:
2JPW - PubMed Abstract:
We present here the solution structure for the bisphosphorylated form of the cardiac N-extension of troponin I (cTnI(1-32)), a region for which there are no previous high-resolution data. Using this structure, the X-ray crystal structure of the cardiac troponin core, and uniform density models of the troponin components derived from neutron contrast variation data, we built atomic models for troponin that show the conformational transition in cardiac troponin induced by bisphosphorylation. In the absence of phosphorylation, our NMR data and sequence analyses indicate a less structured cardiac N-extension with a propensity for a helical region surrounding the phosphorylation motif, followed by a helical C-terminal region (residues 25-30). In this conformation, TnI(1-32) interacts with the N-lobe of cardiac troponin C (cTnC) and thus is positioned to modulate myofilament Ca2+-sensitivity. Bisphosphorylation at Ser23/24 extends the C-terminal helix (residues 21-30) which results in weakening interactions with the N-lobe of cTnC and a re-positioning of the acidic amino terminus of cTnI(1-32) for favorable interactions with basic regions, likely the inhibitory region of cTnI. An extended poly(L-proline)II helix between residues 11 and 19 serves as the rigid linker that aids in re-positioning the amino terminus of cTnI(1-32) upon bisphosphorylation at Ser23/24. We propose that it is these electrostatic interactions between the acidic amino terminus of cTnI(1-32) and the basic inhibitory region of troponin I that induces a bending of cTnI at the end that interacts with cTnC. This model provides a molecular mechanism for the observed changes in cross-bridge kinetics upon TnI phosphorylation.
Organizational Affiliation:
Department of Molecular Genetics, Biochemistry, and Microbiology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, Cincinnati, Ohio, 45267, USA.