Solution structure of the N-domain of Wilson disease protein: Distinct nucleotide-binding environment and effects of disease mutations
Dmitriev, O., Tsivkovskii, R., Abildgaard, F., Morgan, C.T., Markley, J.L., Lutsenko, S.(2006) Proc Natl Acad Sci U S A 103: 5302-5307
- PubMed: 16567646
- DOI: https://doi.org/10.1073/pnas.0507416103
- Primary Citation of Related Structures:
2ARF - PubMed Abstract:
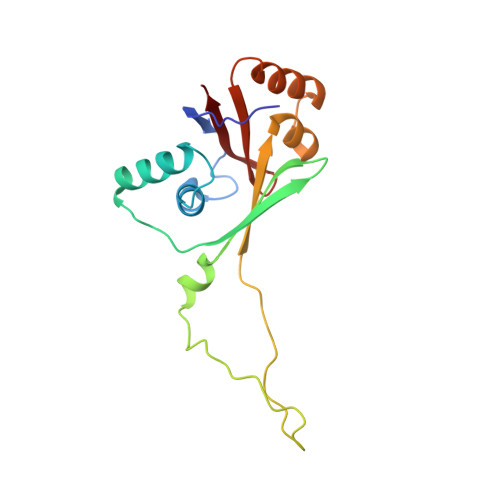
Wilson disease protein (ATP7B) is a copper-transporting P(1B)-type ATPase that regulates copper homeostasis and biosynthesis of copper-containing enzymes in human tissues. Inactivation of ATP7B or related ATP7A leads to severe neurodegenerative disorders, whereas their overexpression contributes to cancer cell resistance to chemotherapeutics. Copper-transporting ATPases differ from other P-type ATPases in their topology and the sequence of their nucleotide-binding domain (N-domain). To gain insight into the structural basis of ATP7B function, we have solved the structure of the ATP7B N-domain in the presence of ATP by using heteronuclear multidimensional NMR spectroscopy. The N-domain consists of a six-stranded beta-sheet with two adjacent alpha-helical hairpins and, unexpectedly, shows higher similarity to the bacterial K(+)-transporting ATPase KdpB than to the mammalian Ca(2+)-ATPase or Na(+),K(+)-ATPase. The common core structure of P-type ATPases is retained in the 3D fold of the N-domain; however, the nucleotide coordination environment of ATP7B within this fold is different. The residues H1069, G1099, G1101, I1102, G1149, and N1150 conserved in the P(1B)-ATPase subfamily contribute to ATP binding. Analysis of the frequent disease mutation H1069Q demonstrates that this mutation does not significantly affect the structure of the N-domain but prevents tight binding of ATP. The structure of the N-domain accounts for the disruptive effects of >30 known Wilson disease mutations. The unique features of the N-domain provide a structural basis for the development of specific inhibitors and regulators of ATP7B.
Organizational Affiliation:
Department of Biomolecular Chemistry, University of Wisconsin, Madison, WI 53706, USA. dmitriev@usask.ca