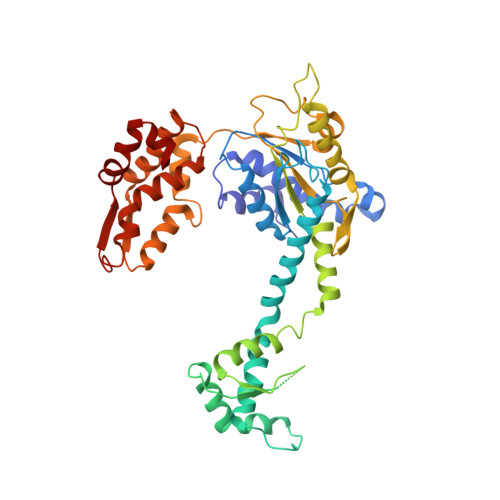
The structures of HsIU and the ATP-dependent protease HsIU-HsIV.
Bochtler, M., Hartmann, C., Song, H.K., Bourenkov, G.P., Bartunik, H.D., Huber, R.(2000) Nature 403: 800-805
- PubMed: 10693812
- DOI: https://doi.org/10.1038/35001629
- Primary Citation of Related Structures:
1DO0, 1DO2 - PubMed Abstract:
The degradation of cytoplasmic proteins is an ATP-dependent process. Substrates are targeted to a single soluble protease, the 26S proteasome, in eukaryotes and to a number of unrelated proteases in prokaryotes. A surprising link emerged with the discovery of the ATP-dependent protease HslVU (heat shock locus VU) in Escherichia coli. Its protease component HslV shares approximately 20% sequence similarity and a conserved fold with 20S proteasome beta-subunits. HslU is a member of the Hsp100 (Clp) family of ATPases. Here we report the crystal structures of free HslU and an 820,000 relative molecular mass complex of HslU and HslV-the first structure of a complete set of components of an ATP-dependent protease. HslV and HslU display sixfold symmetry, ruling out mechanisms of protease activation that require a symmetry mismatch between the two components. Instead, there is conformational flexibility and domain motion in HslU and a localized order-disorder transition in HslV. Individual subunits of HslU contain two globular domains in relative orientations that correlate with nucleotide bound and unbound states. They are surprisingly similar to their counterparts in N-ethylmaleimide-sensitive fusion protein, the prototype of an AAA-ATPase. A third, mostly alpha-helical domain in HslU mediates the contact with HslV and may be the structural equivalent of the amino-terminal domains in proteasomal AAA-ATPases.
Organizational Affiliation:
Max-Planck-Institut für Biochemie, Planegg, Germany.