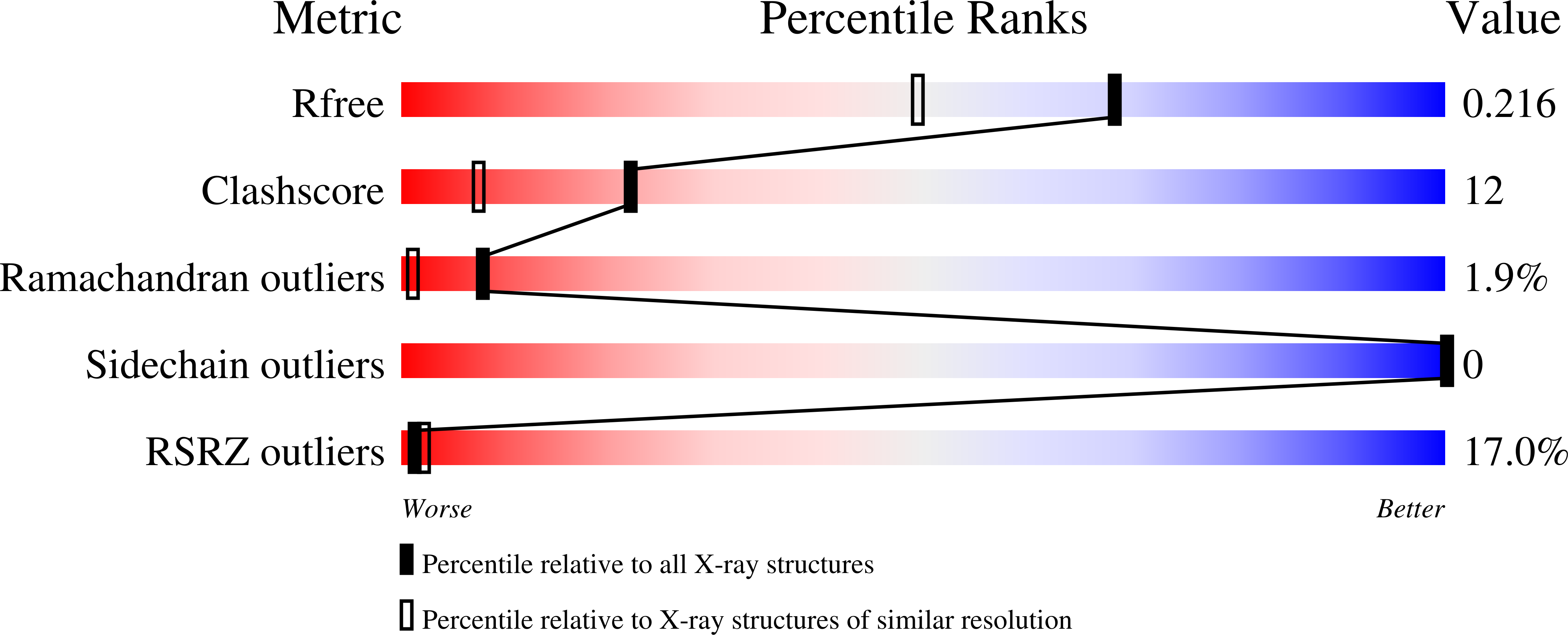
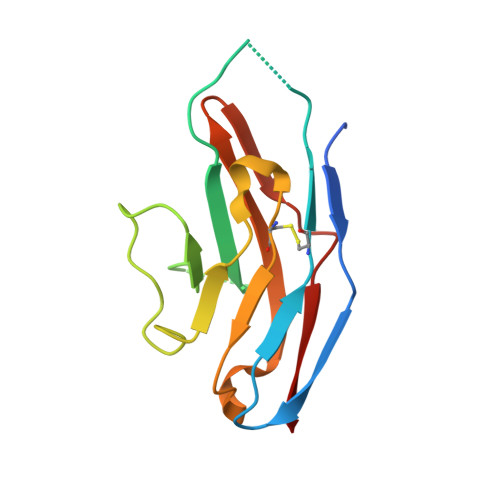
High Resolution Structures of Siglec-7 - Insights Into Ligand Specificity in the Siglec Family
Alphey, M.S., Attrill, H., Crocker, P.R., Van Aalten, D.M.F.(2003) J Biol Chem 278: 3372
- PubMed: 12438315
- DOI: https://doi.org/10.1074/jbc.M210602200
- Primary Citation of Related Structures:
1O7S - PubMed Abstract:
Sialic acid-binding immunoglobulin-like lectins (Siglecs) recognize sialylated glycoconjugates and play a role in cell-cell recognition. Siglec-7 is expressed on natural killer cells and displays unique ligand binding properties different from other members of the Siglec family. Here we describe the high resolution structures of the N-terminal V-set Ig-like domain of Siglec-7 in two crystal forms, at 1.75 and 1.9 A. The latter crystal form reveals the full structure of this domain and allows us to speculate on the differential ligand binding properties displayed by members of the Siglec family. A fully ordered N-linked glycan is observed, tethered by tight interactions with symmetry-related protein molecules in the crystal. Comparison of the structure with that of sialoadhesin and a model of Siglec-9 shows that the unique preference of Siglec-7 for alpha(2,8)-linked disialic acid is likely to reside in the C-C' loop, which is variable in the Siglec family. In the Siglec-7 structure, the ligand-binding pocket is occupied by a loop of a symmetry-related molecule, mimicking the interactions with sialic acid.
Organizational Affiliation:
Division of Cell Biology and Immunology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland, United Kingdom.