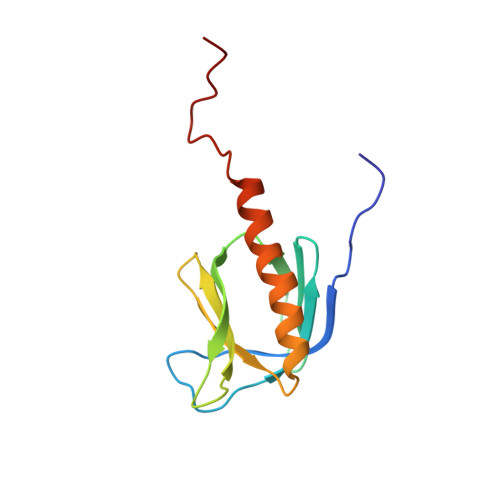
The solution structure and dynamics of the pleckstrin homology domain of G protein-coupled receptor kinase 2 (beta-adrenergic receptor kinase 1). A binding partner of Gbetagamma subunits.
Fushman, D., Najmabadi-Haske, T., Cahill, S., Zheng, J., LeVine 3rd., H., Cowburn, D.(1998) J Biol Chem 273: 2835-2843
- PubMed: 9446593
- DOI: https://doi.org/10.1074/jbc.273.5.2835
- Primary Citation of Related Structures:
1BAK - PubMed Abstract:
The solution structure of an extended pleckstrin homology (PH) domain from the beta-adrenergic receptor kinase is obtained by high resolution NMR. The structure establishes that the beta-adrenergic receptor kinase extended PH domain has the same fold and topology as other PH domains, and there are several unique features, most notably an extended C-terminal alpha-helix that behaves as a molten helix, and a surface charge polarity that is extensively modified by positive residues in the extended alpha-helix and the C terminus. These observations complement biochemical evidence that the C-terminal portion of this PH domain participates in protein-protein interactions with Gbetagamma subunits. This suggests that the C-terminal segment of the PH domain may function to mediate protein-protein interactions with the targets of PH domains.
Organizational Affiliation:
Laboratory of Physical Biochemistry, The Rockefeller University, New York, New York 10021-6399, USA.