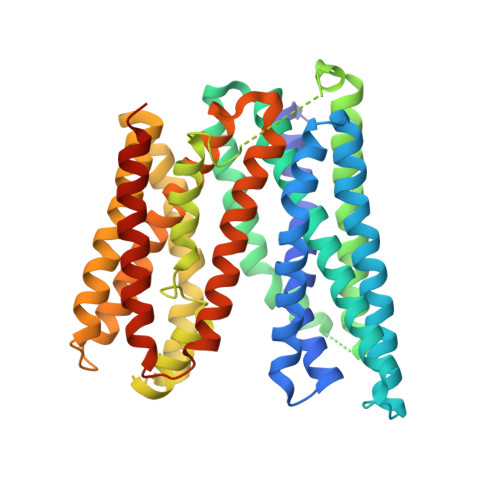
Outward- and inward-facing structures of a putative bacterial transition-metal transporter with homology to ferroportin
Taniguchi, R., Kato, H.E., Font, J., Deshpande, C.N., Wada, M., Ito, K., Ishitani, R., Jormakka, M., Nureki, O.(2015) Nat Commun 6: 8545-8545
- PubMed: 26461048
- DOI: https://doi.org/10.1038/ncomms9545
- Primary Citation of Related Structures:
5AYM, 5AYN, 5AYO - PubMed Abstract:
In vertebrates, the iron exporter ferroportin releases Fe(2+) from cells into plasma, thereby maintaining iron homeostasis. The transport activity of ferroportin is suppressed by the peptide hormone hepcidin, which exhibits upregulated expression in chronic inflammation, causing iron-restrictive anaemia. However, due to the lack of structural information about ferroportin, the mechanisms of its iron transport and hepcidin-mediated regulation remain largely elusive. Here we report the crystal structures of a putative bacterial homologue of ferroportin, BbFPN, in both the outward- and inward-facing states. Despite undetectable sequence similarity, BbFPN adopts the major facilitator superfamily fold. A comparison of the two structures reveals that BbFPN undergoes an intra-domain conformational rearrangement during the transport cycle. We identify a substrate metal-binding site, based on structural and mutational analyses. Furthermore, the BbFPN structures suggest that a predicted hepcidin-binding site of ferroportin is located within its central cavity. Thus, BbFPN may be a valuable structural model for iron homeostasis regulation by ferroportin.
Organizational Affiliation:
Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, Tokyo 113-0032, Japan.