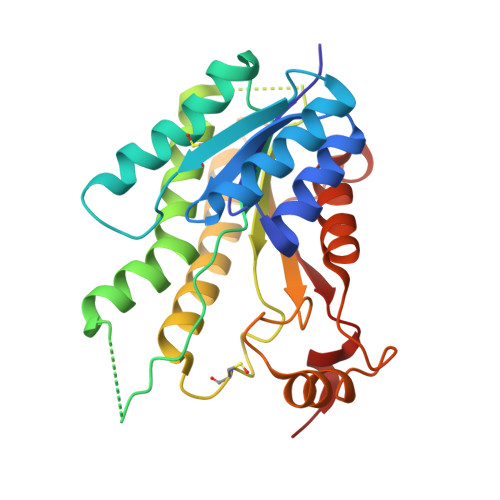
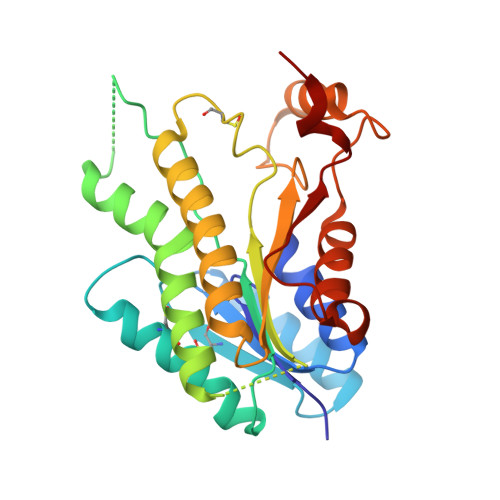
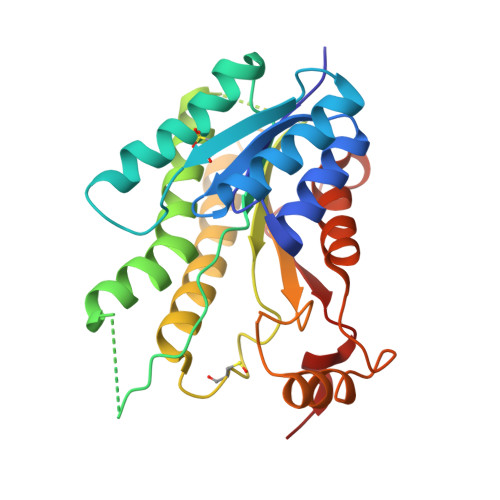
Exploiting the 2-Amino-1,3,4-thiadiazole Scaffold To Inhibit Trypanosoma brucei Pteridine Reductase in Support of Early-Stage Drug Discovery.
Linciano, P., Dawson, A., Pohner, I., Costa, D.M., Sa, M.S., Cordeiro-da-Silva, A., Luciani, R., Gul, S., Witt, G., Ellinger, B., Kuzikov, M., Gribbon, P., Reinshagen, J., Wolf, M., Behrens, B., Hannaert, V., Michels, P.A.M., Nerini, E., Pozzi, C., di Pisa, F., Landi, G., Santarem, N., Ferrari, S., Saxena, P., Lazzari, S., Cannazza, G., Freitas-Junior, L.H., Moraes, C.B., Pascoalino, B.S., Alcantara, L.M., Bertolacini, C.P., Fontana, V., Wittig, U., Muller, W., Wade, R.C., Hunter, W.N., Mangani, S., Costantino, L., Costi, M.P.(2017) ACS Omega 2: 5666-5683
- PubMed: 28983525
- DOI: https://doi.org/10.1021/acsomega.7b00473
- Primary Citation of Related Structures:
2YHI, 2YHU, 4WCD, 4WCF, 5IZC - PubMed Abstract:
Pteridine reductase-1 (PTR1) is a promising drug target for the treatment of trypanosomiasis. We investigated the potential of a previously identified class of thiadiazole inhibitors of Leishmania major PTR1 for activity against Trypanosoma brucei ( Tb ). We solved crystal structures of several Tb PTR1-inhibitor complexes to guide the structure-based design of new thiadiazole derivatives. Subsequent synthesis and enzyme- and cell-based assays confirm new, mid-micromolar inhibitors of Tb PTR1 with low toxicity. In particular, compound 4m , a biphenyl-thiadiazole-2,5-diamine with IC 50 = 16 μM, was able to potentiate the antitrypanosomal activity of the dihydrofolate reductase inhibitor methotrexate (MTX) with a 4.1-fold decrease of the EC 50 value. In addition, the antiparasitic activity of the combination of 4m and MTX was reversed by addition of folic acid. By adopting an efficient hit discovery platform, we demonstrate, using the 2-amino-1,3,4-thiadiazole scaffold, how a promising tool for the development of anti- T. brucei agents can be obtained.
Organizational Affiliation:
Dipartimento di Scienze della Vita, Università degli Studi di Modena e Reggio Emilia, Via Campi 103, 41125 Modena, Italy.