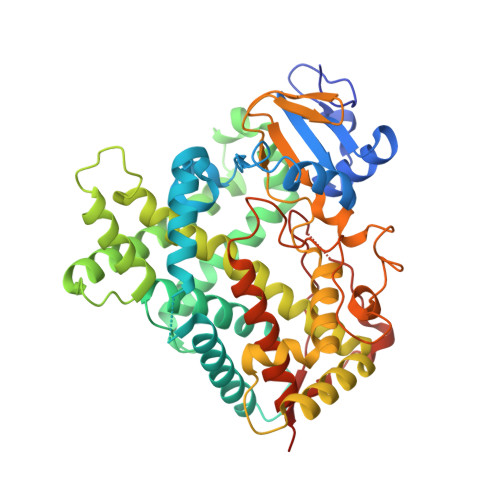
A Structural Snapshot of CYP2B4 in Complex with Paroxetine Provides Insights into Ligand Binding and Clusters of Conformational States.
Shah, M.B., Kufareva, I., Pascual, J., Zhang, Q., Stout, C.D., Halpert, J.R.(2013) J Pharmacol Exp Ther 346: 113-120
- PubMed: 23633618
- DOI: https://doi.org/10.1124/jpet.113.204776
- Primary Citation of Related Structures:
4JLT - PubMed Abstract:
An X-ray crystal structure of CYP2B4 in complex with the drug paroxetine [(3S,4R)-3-[(2H-1,3-benzodioxol-5-yloxy)methyl]-4-(4-fluorophenyl)piperidine] was solved at 2.14 Å resolution. The structure revealed a conformation intermediate to that of the recently solved complex with amlodipine and that of the more compact complex with 4-(4-chlorophenyl)imidazole in terms of the placement of the F-G cassette. Moreover, comparison of the new structure with 15 previously solved structures of CYP2B4 revealed some new insights into the determinants of active-site size and shape. The 2B4-paroxetine structure is nearly superimposable on a previously solved closed structure in a ligand-free state. Despite the overall conformational similarity among multiple closed structures, the active-site cavity volume of the paroxetine complex is enlarged. Further analysis of the accessible space and binding pocket near the heme reveals a new subchamber that resulted from the movement of secondary structural elements and rearrangements of active-site side chains. Overall, the results from the comparison of all 16 structures of CYP2B4 demonstrate a cluster of protein conformations that were observed in the presence or absence of various ligands.
Organizational Affiliation:
Skaggs School of Pharmacy and Pharmaceutical Sciences, University of California San Diego, 9500 Gilman Drive, Mail Code 0703, La Jolla, CA 92093-0703, USA. m7shah@ucsd.edu