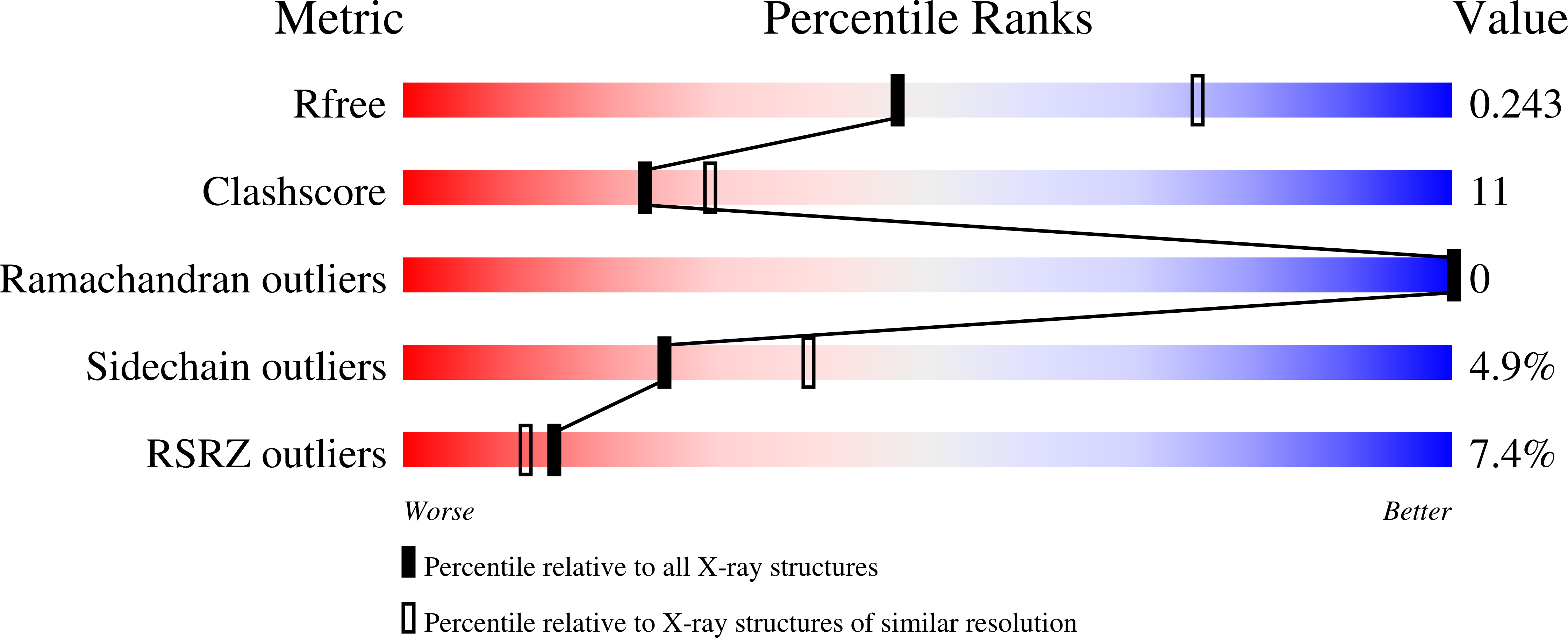
Bacillus anthracis inosine 5'-monophosphate dehydrogenase in action: the first bacterial series of structures of phosphate ion-, substrate-, and product-bound complexes.
Makowska-Grzyska, M., Kim, Y., Wu, R., Wilton, R., Gollapalli, D.R., Wang, X.K., Zhang, R., Jedrzejczak, R., Mack, J.C., Maltseva, N., Mulligan, R., Binkowski, T.A., Gornicki, P., Kuhn, M.L., Anderson, W.F., Hedstrom, L., Joachimiak, A.(2012) Biochemistry 51: 6148-6163
- PubMed: 22788966
- DOI: https://doi.org/10.1021/bi300511w
- Primary Citation of Related Structures:
3TSB, 3TSD, 3USB - PubMed Abstract:
Inosine 5'-monophosphate dehydrogenase (IMPDH) catalyzes the first unique step of the GMP branch of the purine nucleotide biosynthetic pathway. This enzyme is found in organisms of all three kingdoms. IMPDH inhibitors have broad clinical applications in cancer treatment, as antiviral drugs and as immunosuppressants, and have also displayed antibiotic activity. We have determined three crystal structures of Bacillus anthracis IMPDH, in a phosphate ion-bound (termed "apo") form and in complex with its substrate, inosine 5'-monophosphate (IMP), and product, xanthosine 5'-monophosphate (XMP). This is the first example of a bacterial IMPDH in more than one state from the same organism. Furthermore, for the first time for a prokaryotic enzyme, the entire active site flap, containing the conserved Arg-Tyr dyad, is clearly visible in the structure of the apoenzyme. Kinetic parameters for the enzymatic reaction were also determined, and the inhibitory effect of XMP and mycophenolic acid (MPA) has been studied. In addition, the inhibitory potential of two known Cryptosporidium parvum IMPDH inhibitors was examined for the B. anthracis enzyme and compared with those of three bacterial IMPDHs from Campylobacter jejuni, Clostridium perfringens, and Vibrio cholerae. The structures contribute to the characterization of the active site and design of inhibitors that specifically target B. anthracis and other microbial IMPDH enzymes.
Organizational Affiliation:
Center for Structural Genomics of Infectious Diseases, University of Chicago, 5735 South Ellis Avenue, Chicago, IL 60637, USA.