The Structural Basis for Peptidomimetic Inhibition of Eukaryotic Ribonucleotide Reductase: A Conformationally Flexible Pharmacophore
Xu, H., Fairman, J.W., Wijerathna, S.R., Kreischer, N.R., LaMacchia, J., Helmbrecht, E., Cooperman, B.S., Dealwis, C.(2008) J Med Chem 51: 4653-4659
- PubMed: 18610997
- DOI: https://doi.org/10.1021/jm800350u
- Primary Citation of Related Structures:
2ZLF, 2ZLG - PubMed Abstract:
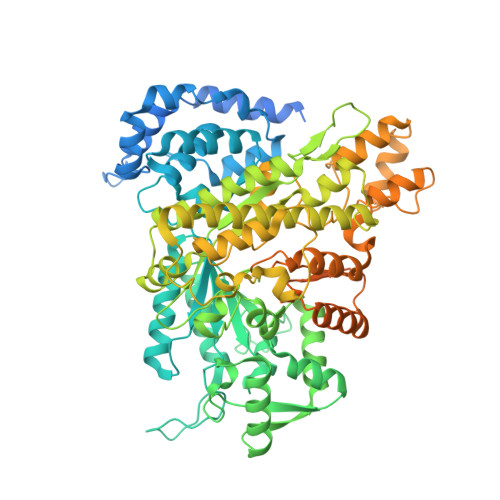
Eukaryotic ribonucleotide reductase (RR) catalyzes nucleoside diphosphate conversion to deoxynucleoside diphosphate. Crucial for rapidly dividing cells, RR is a target for cancer therapy. RR activity requires formation of a complex between subunits R1 and R2 in which the R2 C-terminal peptide binds to R1. Here we report crystal structures of heterocomplexes containing mammalian R2 C-terminal heptapeptide, P7 (Ac-1FTLDADF7) and its peptidomimetic P6 (1Fmoc(Me)PhgLDChaDF7) bound to Saccharomyces cerevisiae R1 (ScR1). P7 and P6, both of which inhibit ScRR, each bind at two contiguous sites containing residues that are highly conserved among eukaryotes. Such binding is quite distinct from that reported for prokaryotes. The Fmoc group in P6 peptide makes several hydrophobic interactions that contribute to its enhanced potency in binding to ScR1. Combining all of our results, we observe three distinct conformations for peptide binding to ScR1. These structures provide pharmacophores for designing highly potent nonpeptide class I RR inhibitors.
Organizational Affiliation:
Department of Pharmacology, Case School of Medicine, Case Western Reserve University, 10900 Euclid Avenue, Cleveland, Ohio 44106-4965, USA.