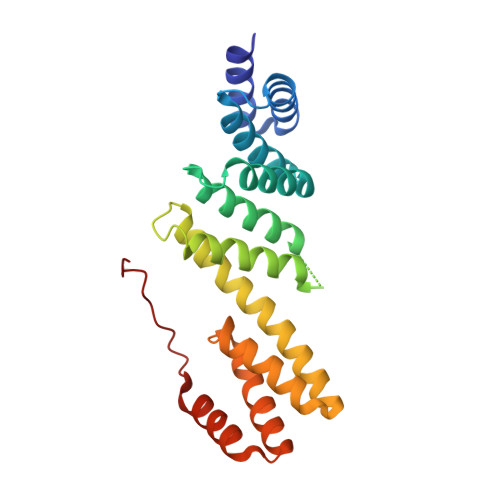
Structural Basis of Outer Membrane Protein Biogenesis in Bacteria.
Albrecht, R., Zeth, K.(2011) J Biol Chem 286: 27792
- PubMed: 21586578
- DOI: https://doi.org/10.1074/jbc.M111.238931
- Primary Citation of Related Structures:
2YH3, 2YH5, 2YH6, 2YH9, 2YHC - PubMed Abstract:
In Escherichia coli, a multicomponent BAM (β-barrel assembly machinery) complex is responsible for recognition and assembly of outer membrane β-barrel proteins. The functionality of BAM in protein biogenesis is mainly orchestrated through the presence of two essential components, BamA and BamD. Here, we present crystal structures of four lipoproteins (BamB-E). Monomeric BamB and BamD proteins display scaffold architectures typically implied in transient protein interactions. BamB is a β-propeller protein comprising eight WD40 repeats. BamD shows an elongated fold on the basis of five tetratricopeptide repeats, three of which form the scaffold for protein recognition. The rod-shaped BamC protein has evolved through the gene duplication of two conserved domains known to mediate protein interactions in structurally related complexes. By contrast, the dimeric BamE is formed through a domain swap and indicates fold similarity to the β-lactamase inhibitor protein family, possibly integrating cell wall stability in BAM function. Structural and biochemical data show evidence for the specific recognition of amphipathic sequences through the tetratricopeptide repeat architecture of BamD. Collectively, our data advance the understanding of the BAM complex and highlight the functional importance of BamD in amphipathic outer membrane β-barrel protein motif recognition and protein delivery.
Organizational Affiliation:
Department of Protein Evolution, Max Planck Institute for Developmental Biology, Spemannstrasse 35, Tübingen 72076, Germany.