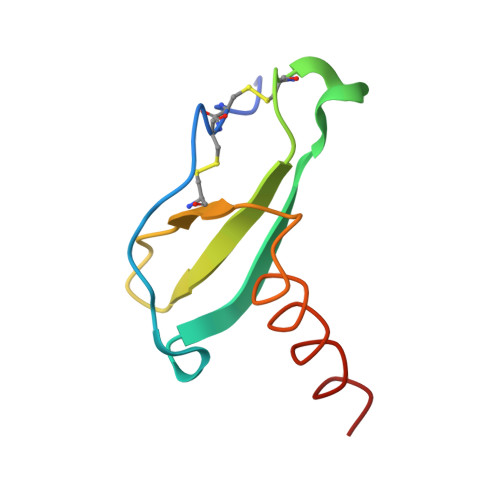
Structure of human MIP-3alpha chemokine.
Malik, Z.A., Tack, B.F.(2006) Acta Crystallogr Sect F Struct Biol Cryst Commun 62: 631-634
- PubMed: 16820679
- DOI: https://doi.org/10.1107/S1744309106006890
- Primary Citation of Related Structures:
2HCI - PubMed Abstract:
The structure of the human macrophage inflammatory protein-3alpha (MIP-3alpha) has been determined at 1.81 angstroms resolution by X-ray crystallography. The dimer crystallized in the tetragonal space group I4, with unit-cell parameters a = b = 83.99, c = 57.20 angstroms. The crystals exhibit two molecules in the asymmetric unit. The structure was solved by the molecular-replacement method and the model was refined to a conventional R value of 20.6% (R(free) = 25.7%). MIP-3alpha possesses the same monomeric structure as previously described for other chemokines. However, in addition to limited structural changes in the beta1-beta2 hairpin of monomer B, the electron density is fully defined for a few extra residues at the N- and C-termini of monomer A and the C-terminus of monomer B compared with MIP-3alpha in space group P6(1). As the N-terminal and loop regions have been shown to be critical for receptor binding and signaling, this additional structural information may help in determining the basis of the CCR6 selectivity of MIP-3alpha.
Organizational Affiliation:
Department of Pharmacology, Carver College of Medicine, The University of Iowa, Iowa City, IA 52242, USA. z-malik@uiowa.edu