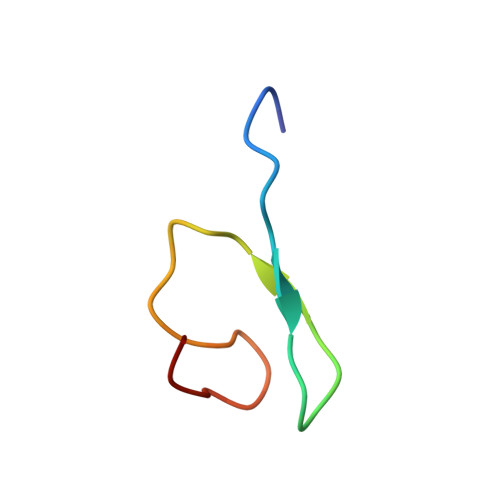
Molecular Characterization of the Ran-binding Zinc Finger Domain of Nup153.
Higa, M.M., Alam, S.L., Sundquist, W.I., Ullman, K.S.(2007) J Biol Chem 282: 17090-17100
- PubMed: 17426026
- DOI: https://doi.org/10.1074/jbc.M702715200
- Primary Citation of Related Structures:
2GQE - PubMed Abstract:
The nuclear pore complex is the gateway for selective traffic between the nucleus and cytoplasm. To learn how building blocks of the pore can create specific docking sites for transport receptors and regulatory factors, we have studied a zinc finger module present in multiple copies within the nuclear pores of higher eukaryotes. All four zinc fingers of human Nup153 were found to bind the small GTPase Ran with dissociation constants ranging between 5 and 40 mum. In addition a fragment of Nup153 encompassing the four tandem zinc fingers was found to bind Ran with similar affinity. NMR structural studies revealed that a representative Nup153 zinc finger adopts the same zinc ribbon structure as the previously characterized Npl4 NZF module. Ran binding was mediated by a three-amino acid motif (Leu(13)/Val(14)/Asn(25)) located within the two zinc coordination loops. Nup153 ZnFs bound GDP and GTP forms of Ran with similar affinities, indicating that this interaction is not influenced by a nucleotide-dependent conformational switch. Taken together, these studies elucidate the Ran-binding interface on Nup153 and, more broadly, provide insight into the versatility of this zinc finger binding module.
Organizational Affiliation:
Department of Oncological Sciences, Huntsman Cancer Institute, University of Utah, Salt Lake City, Utah 84112, USA.