Somatostatin receptor 1 selective analogues: 4. Three-dimensional consensus structure by NMR
Grace, C.R.R., Durrer, L., Koerber, S.C., Erchegyi, J., Reubi, J.C., Rivier, J.E., Riek, R.(2005) J Med Chem 48: 523-533
- PubMed: 15658866
- DOI: https://doi.org/10.1021/jm049518u
- Primary Citation of Related Structures:
1XXZ, 1XY4, 1XY5, 1XY6, 1XY8, 1XY9 - PubMed Abstract:
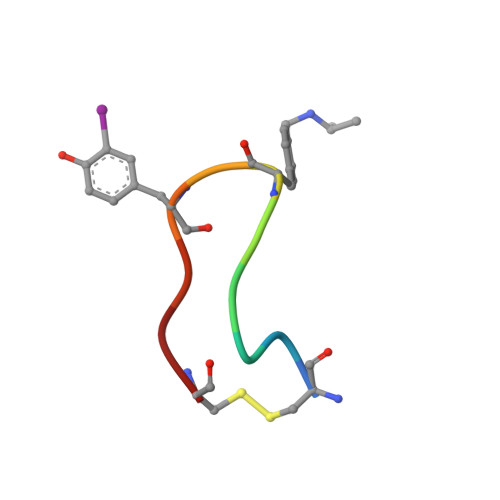
The three-dimensional NMR structures of six analogues of somatostatin (SRIF) are described. These analogues with the amino acid 4-(N-isopropyl)-aminomethylphenylalanine (IAmp) at position 9 exhibit potent and highly selective binding to human SRIF subtype 1 receptors (sst(1)). The conformations reveal that the backbones of these analogues have a hairpin-like structure similar to the sst(2)-subtype-selective analogues. This structure serves as a scaffold for retaining a unique arrangement of the side chains of d-Trp(8), IAmp(9), Phe(7), and Phe(11) or m-I-Tyr(11) (m-I-Tyr = mono-iodo-tyrosine). The conformational preferences and results from biological analyses of these analogues(1,2) allow a detailed study of the structure-activity relationship of SRIF. The proposed consensus pharmacophore of the sst(1)-selective analogues requires a unique set of distances between an indole/2-naphthyl ring, an IAmp side chain, and two aromatic rings. This motif is necessary and sufficient to explain the binding affinities of all of the analogues studied and is distinct from the existing models suggested for sst(4) as well as sst(2)/sst(5) selectivity.
Organizational Affiliation:
Structural Biology Laboratory, The Salk Institute for Biological Studies, 10010 North Torrey Pines Road, La Jolla, California 92037, USA.