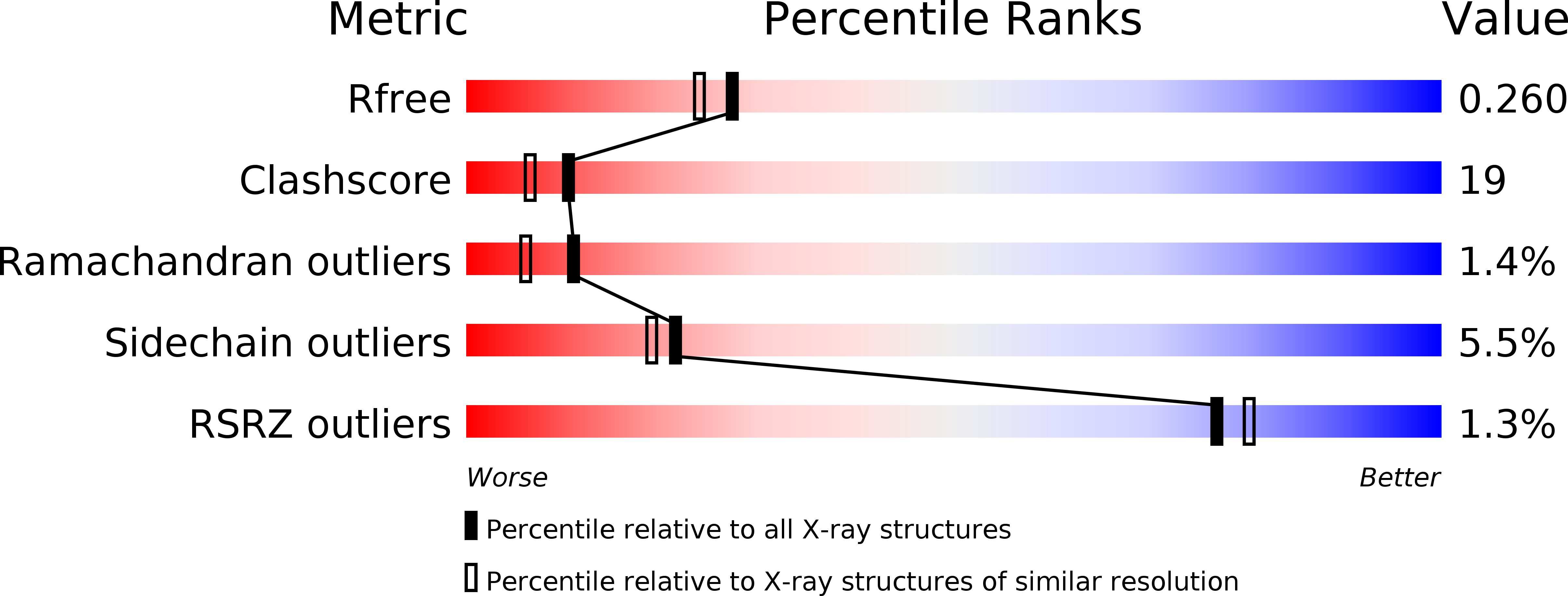
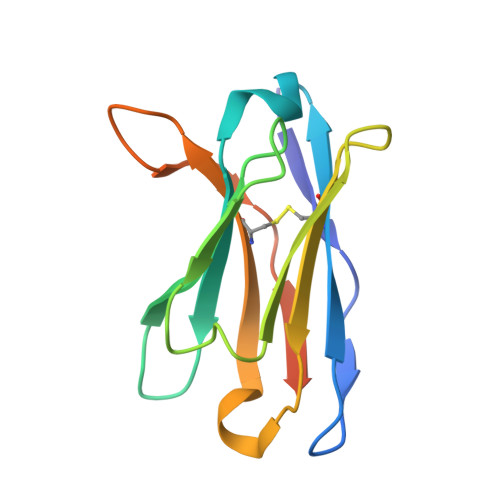
Thermodynamic consequences of mutations in vernier zone residues of a humanized anti-human epidermal growth factor receptor murine antibody, 528
Makabe, K., Nakanishi, T., Tsumoto, K., Tanaka, Y., Kondo, H., Umetsu, M., Sone, Y., Asano, R., Kumagai, I.(2008) J Biol Chem 283: 1156-1166
- PubMed: 17947238
- DOI: https://doi.org/10.1074/jbc.M706190200
- Primary Citation of Related Structures:
1WT5, 2Z4Q - PubMed Abstract:
To investigate the role of Vernier zone residues, which are comprised in the framework regions and underlie the complementarity-determining regions (CDRs) of antibodies, in the specific, high affinity interactions of antibodies with their targets, we focused on the variable domain fragment of murine anti-human epidermal growth factor receptor antibody 528 (m528Fv). Grafting of the CDRs of m528Fv onto a selected framework region of human antibodies, referred to as humanization, reduced the antibody's affinity for its target by a factor of 1/40. The reduction in affinity was due to a substantial reduction in the negative enthalpy change associated with binding. Crystal structures of the ligand-free antibody fragments showed no noteworthy conformational changes due to humanization, and the loop structures of the CDRs of the humanized antibodies were identical to those of the parent antibodies. Several mutants of the CDR-grafted (humanized) variable domain fragment (h528Fv), in which some of the Vernier zone residues in the heavy chain were replaced with the parental murine residues, were constructed and prepared using a bacterial expression system. Thermodynamic analyses of the interactions between the mutants and the soluble extracellular domain of epidermal growth factor receptor showed that several single mutations and a double mutation increased the negative enthalpy and heat capacity changes. Combination of these mutations, however, led to somewhat reduced negative enthalpy and heat capacity changes. The affinity of each mutant for the target was within the range for the wild-type h528Fv, and this similarity was due to enthalpy-entropy compensation. These results suggest that Vernier zone residues make enthalpic contributions to antigen binding and that the regulation of conformational entropy changes upon humanization of murine antibodies must be carefully considered and optimized.
Organizational Affiliation:
Department of Biomolecular Engineering, Graduate School of Engineering, Tohoku University, Aoba-yama 6-6-11-606, Aoba-ku, Sendai 980-8579, Japan.