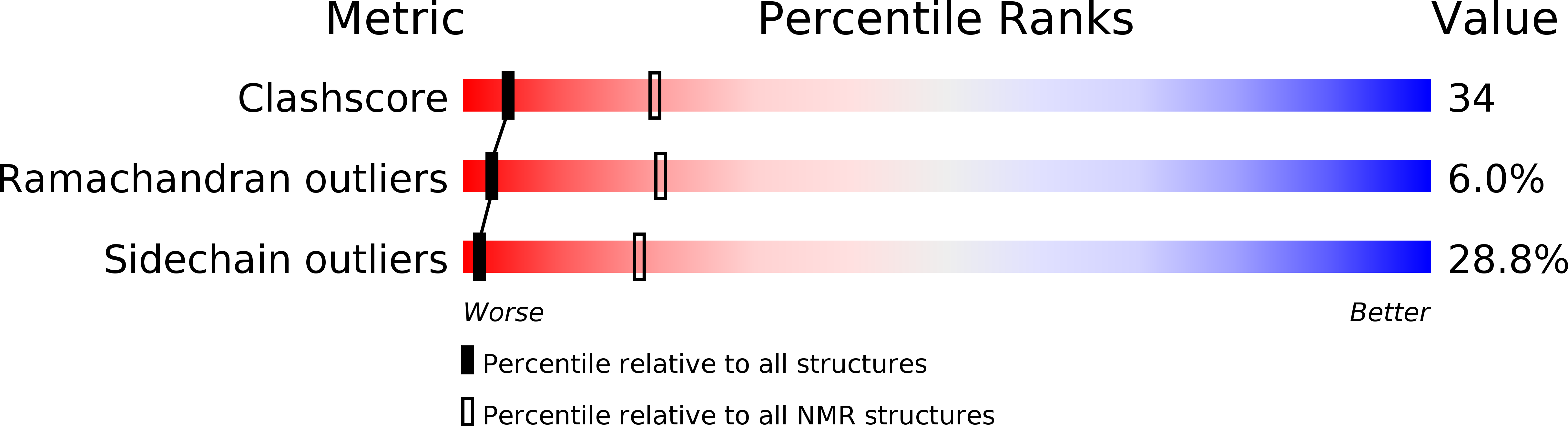Solution structure of the kringle domain from urokinase-type plasminogen activator.
Li, X., Bokman, A.M., Llinas, M., Smith, R.A., Dobson, C.M.(1994) J Mol Biol 235: 1548-1559
- PubMed: 8107091
- DOI: https://doi.org/10.1006/jmbi.1994.1106
- Primary Citation of Related Structures:
1KDU - PubMed Abstract:
The solution structure of the kringle domain from urokinase-type plasminogen activator (u-PA) has been determined using 1H nuclear magnetic resonance spectroscopy and dynamical simulated annealing calculations. A total of 35 structures, 20 generated using a distance geometry method prior to simulated annealing and 15 generated using initial random phi, psi values, have been calculated based on 946 experimental nuclear Overhauser effect distance constraints and 48 dihedral angle constraints. Excluding the N- and C-terminal residues (-1 to 12, 77 to 82) and a number of surface residues (M18, G19, S42, D55 to R60, G67) that are disordered or flexible, the root mean square deviation values from the mean structure are 0.49(+/- 0.14) A and 0.65(+/- 0.16) A for the backbone atoms, and 1.03(+/- 0.21) A and 1.39(+/- 0.24) A for all heavy atoms, for the two sets of structures, respectively. An extended binding site for anionic polysaccharides such as heparin has been located on a relatively flat facet of the molecule, involving three consecutive arginines, R57, R58 and R60 (there is a deletion at site 59 of the consensus sequence), which form a cationic triad facing the solvent, and two histidines, H37 and H40, at the opposite end of the molecule. Comparison between the u-PA kringle structure and the crystal and NMR solution structures of tissue-type plasminogen activator kringle 2 has shown that the two proteins have similar global folds but demonstrate a number of local differences.
Organizational Affiliation:
Oxford Centre for Molecular Sciences, University of Oxford, U.K.