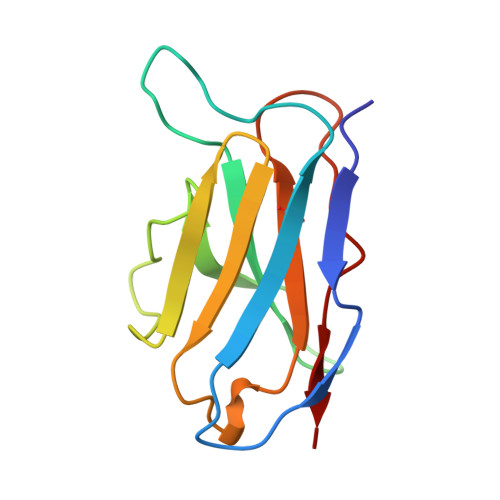
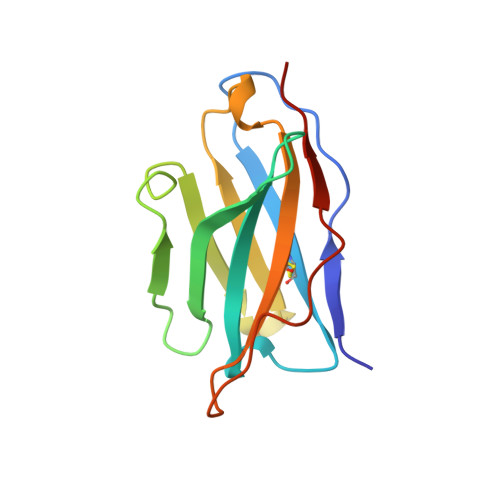
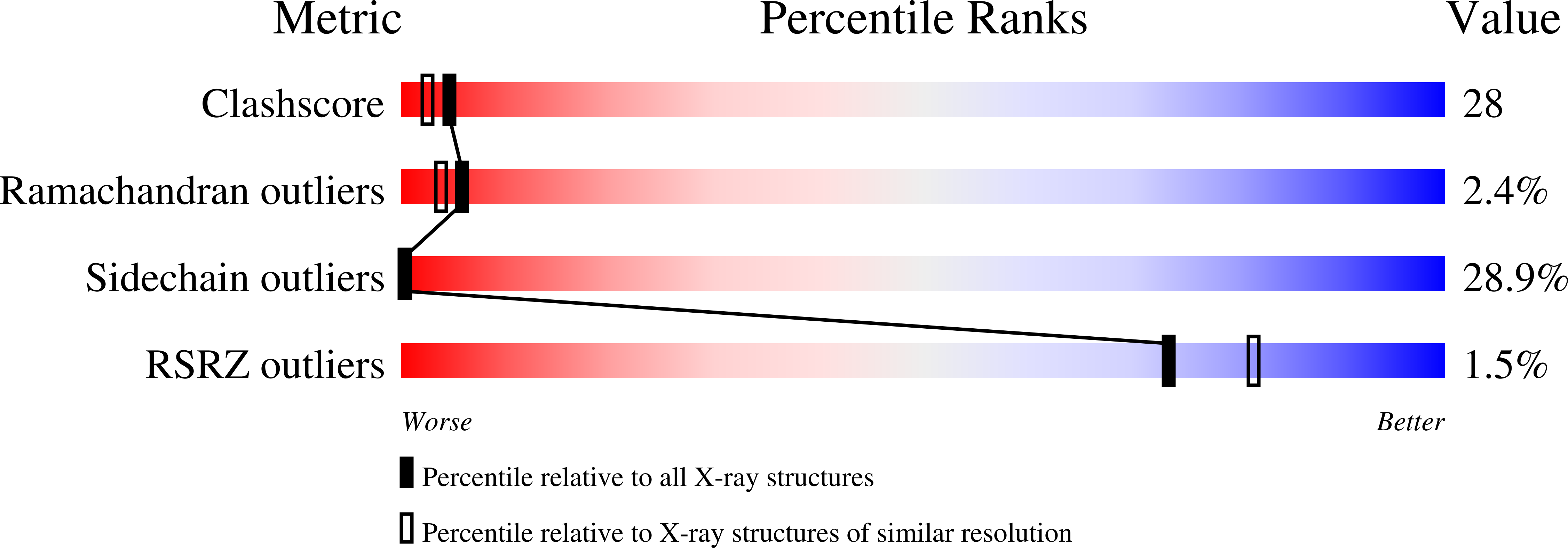A single chain Fv fragment of P-glycoprotein-specific monoclonal antibody C219. Design, expression, and crystal structure at 2.4 A resolution.
Hoedemaeker, F.J., Signorelli, T., Johns, K., Kuntz, D.A., Rose, D.R.(1997) J Biol Chem 272: 29784-29789
- PubMed: 9368049
- DOI: https://doi.org/10.1074/jbc.272.47.29784
- Primary Citation of Related Structures:
1AP2 - PubMed Abstract:
A construct encoding a single chain variable fragment of the anti-P-glycoprotein monoclonal antibody C219 was made by combining the coding sequences for the heavy and light chain variable domains with a sequence encoding the flexible linker (GGGGS)3, an OmpA signal sequence, a c-myc identification tag, and a five-histidine purification tag. The construct was expressed in Escherichia coli and purified from the periplasmic fraction using a nickel chelate column and ion exchange chromatography. Three-step Western blot analysis showed that the construct retains binding affinity for P-glycoprotein. Crystals of 1.0 x 0.2 x 0.2 mm were grown in 100 mM citrate, pH 4.5, 21% polyethylene glycol 6000 in the presence of low concentrations of subtilisin, resulting in proteolytic removal of the linker and purification tags. The structure was solved to a resolution of 2.4 A with an R factor of 20.6, an Rfree of 28.5, and good stereochemistry. This result could lead to a clinically useful product based on antibody C219 for the diagnosis of P-glycoprotein-mediated multidrug resistance. The molecule will also be useful in biophysical studies of functional domains of P-glycoprotein, as well as studies of the intact molecule.
Organizational Affiliation:
Ontario Cancer Institute and Department of Medical Biophysics, University of Toronto, Toronto M5G 2M9, Ontario, Canada.