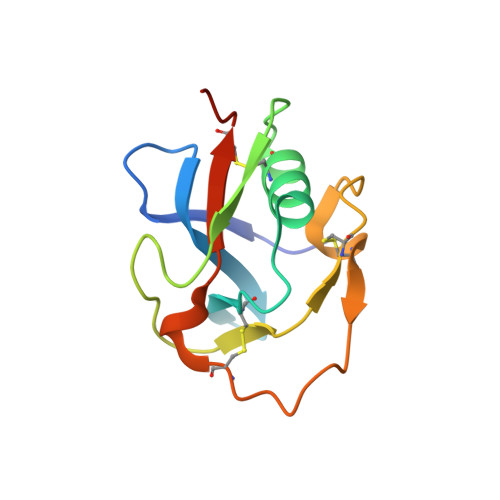
Structural characterization of the third scavenger receptor cysteine-rich domain of murine neurotrypsin.
Canciani, A., Catucci, G., Forneris, F.(2019) Protein Sci 28: 746-755
- PubMed: 30748049
- DOI: https://doi.org/10.1002/pro.3587
- Primary Citation of Related Structures:
6H8M - PubMed Abstract:
Neurotrypsin (NT) is a multi-domain serine protease of the nervous system with only one known substrate: the large proteoglycan Agrin. NT has seen to be involved in the maintenance/turnover of neuromuscular junctions and in processes of synaptic plasticity in the central nervous system. Roles which have been tied to its enzymatic activity, localized in the C-terminal serine-protease (SP) domain. However the purpose of NT's remaining 3-4 scavenger receptor cysteine-rich (SRCR) domains is still unclear. We have determined the crystal structure of the third SRCR domain of murine NT (mmNT-SRCR3), immediately preceding the SP domain and performed a comparative structural analysis using homologous SRCR structures. Our data and the elevated degree of structural conservation with homologous domains highlight possible functional roles for NT SRCRs. Computational and experimental analyses suggest the identification of a putative binding region for Ca 2+ ions, known to regulate NT enzymatic activity. Furthermore, sequence and structure comparisons allow to single out regions of interest that, in future studies, might be implicated in Agrin recognition/binding or in interactions with as of yet undiscovered NT partners.
Organizational Affiliation:
The Armenise-Harvard Laboratory of Structural Biology, Department of Biology and Biotechnology, University of Pavia, Via Ferrata 9/A, 27100 Pavia, Italy.