Artificial cysteine-lipases with high activity and altered catalytic mechanism created by laboratory evolution.
Cen, Y., Singh, W., Arkin, M., Moody, T.S., Huang, M., Zhou, J., Wu, Q., Reetz, M.T.(2019) Nat Commun 10: 3198-3198
- PubMed: 31324776
- DOI: https://doi.org/10.1038/s41467-019-11155-3
- Primary Citation of Related Structures:
6ISP, 6ISQ, 6ISR - PubMed Abstract:
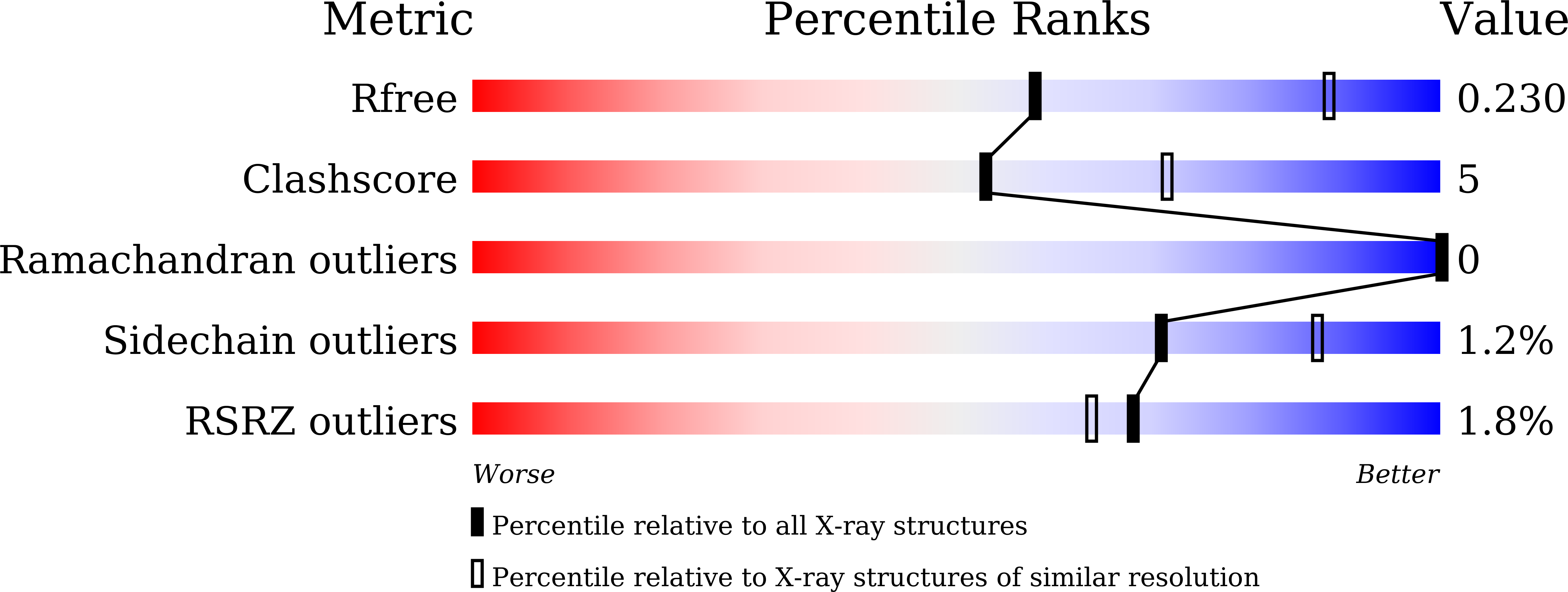
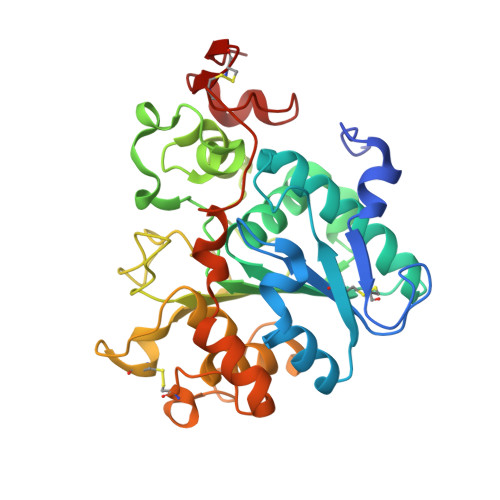
Engineering artificial enzymes with high activity and catalytic mechanism different from naturally occurring enzymes is a challenge in protein design. For example, many attempts have been made to obtain active hydrolases by introducing a Ser → Cys exchange at the respective catalytic triads, but this generally induced a breakdown of activity. We now report that this long-standing dogma no longer pertains, provided additional mutations are introduced by directed evolution. By employing Candida antarctica lipase B (CALB) as the model enzyme with the Ser-His-Asp catalytic triad, a highly active cysteine-lipase having a Cys-His-Asp catalytic triad and additional mutations W104V/A281Y/A282Y/V149G can be evolved, showing a 40-fold higher catalytic efficiency than wild-type CALB in the hydrolysis of 4-nitrophenyl benzoate, and tolerating bulky substrates. Crystal structures, kinetics, MD simulations and QM/MM calculations reveal dynamic features and explain all results, including the preference of a two-step mechanism involving the zwitterionic pair Cys105 - /His224 + rather than a concerted process.
Organizational Affiliation:
Department of Chemistry, Zhejiang University, 310027, Hangzhou, China.